| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected and Quantified |
|---|
| Creation Date | 2005-11-16 15:48:42 UTC |
|---|
| Update Date | 2023-05-30 20:55:56 UTC |
|---|
| HMDB ID | HMDB0000191 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
| Common Name | L-Aspartic acid |
|---|
| Description | Aspartic acid (Asp), also known as L-aspartic acid or as aspartate, the name of its anion, is an alpha-amino acid. These are amino acids in which the amino group is attached to the carbon atom immediately adjacent to the carboxylate group (alpha carbon). Amino acids are organic compounds that contain amino (-NH2) and carboxyl (-COOH) functional groups, along with a side chain (R group) specific to each amino acid. L-aspartic acid is one of 20 proteinogenic amino acids, i.e., the amino acids used in the biosynthesis of proteins. Aspartic acid is found in all organisms ranging from bacteria to plants to animals. It is classified as an acidic, charged (at physiological pH), aliphatic amino acid. In humans, aspartic acid is a nonessential amino acid derived from glutamic acid by enzymes using vitamin B6. However, in the human body, aspartate is most frequently synthesized through the transamination of oxaloacetate. A non-essential amino acid is an amino acid that can be synthesized from central metabolic pathway intermediates in humans and is not required in the diet. As its name indicates, aspartic acid is the carboxylic acid analog of asparagine. The D-isomer of aspartic acid (D-aspartic acid) is one of two D-amino acids commonly found in mammals. Aspartic acid was first discovered in 1827 by Auguste-Arthur Plisson and Étienne Ossian Henry by hydrolysis of asparagine, which had been isolated from asparagus juice in 1806. Aspartate has many biochemical roles. It is a neurotransmitter, a metabolite in the urea cycle and it participates in gluconeogenesis. It carries reducing equivalents in the malate-aspartate shuttle, which utilizes the ready interconversion of aspartate and oxaloacetate, which is the oxidized (dehydrogenated) derivative of malic acid. Aspartate donates one nitrogen atom in the biosynthesis of inosine, the precursor to the purine bases which are key to DNA biosynthesis. In addition, aspartic acid acts as a hydrogen acceptor in a chain of ATP synthase. Aspartic acid is a major excitatory neurotransmitter, which is sometimes found to be increased in epileptic and stroke patients. It is decreased in depressed patients and in patients with brain atrophy. As a neurotransmitter, aspartic acid may provide resistance to fatigue and thus lead to endurance, although the evidence to support this idea is not strong (Wikipedia ). Aspartic acid supplements are being evaluated. Five grams can raise blood levels. Magnesium and zinc may be natural inhibitors of some of the actions of aspartic acid. Aspartic acid, when chemically coupled with the amino acid D-phenylalanine, is a part of a natural sweetener, aspartame. This sweetener is an advance in artificial sweeteners, and is probably safe in normal doses to all except phenylketonurics. Aspartic acid may be a significant immunostimulant of the thymus and can protect against some of the damaging effects of radiation. Aspartic acid is found in higher abundance in: oysters, luncheon meats, sausage meat, wild game, sprouting seeds, oat flakes, avocado, asparagus, young sugarcane, and molasses from sugar beets. | Read more...
|---|
| Structure | InChI=1S/C4H7NO4/c5-2(4(8)9)1-3(6)7/h2H,1,5H2,(H,6,7)(H,8,9)/t2-/m0/s1 |
|---|
| Synonyms | | Value | Source |
|---|
| (S)-2-Aminobutanedioic acid | ChEBI | | (S)-2-Aminosuccinic acid | ChEBI | | 2-Aminosuccinic acid | ChEBI | | Asp | ChEBI | | ASPARTIC ACID | ChEBI | | D | ChEBI | | L-Asparaginsaeure | ChEBI | | L-Asp | Kegg | | (S)-2-Aminobutanedioate | Generator | | (S)-2-Aminosuccinate | Generator | | 2-Aminosuccinate | Generator | | ASPARTate | Generator | | L-Aspartate | Generator | | (+)-Aspartate | HMDB | | (+)-Aspartic acid | HMDB | | (2S)-Aspartate | HMDB | | (2S)-Aspartic acid | HMDB | | (L)-Aspartate | HMDB | | (L)-Aspartic acid | HMDB | | (R)-2-Aminosuccinate | HMDB | | (S)-(+)-Aspartate | HMDB | | (S)-(+)-Aspartic acid | HMDB | | (S)-Amino-butanedioate | HMDB | | (S)-Amino-butanedioic acid | HMDB | | (S)-Aminobutanedioate | HMDB | | (S)-Aminobutanedioic acid | HMDB | | (S)-Aspartate | HMDB | | (S)-Aspartic acid | HMDB | | 2-Amino-3-methylsuccinate | HMDB | | 2-Amino-3-methylsuccinic acid | HMDB | | alpha-Aminosuccinate | HMDB | | alpha-Aminosuccinic acid | HMDB | | Aminosuccinate | HMDB | | Asparagate | HMDB | | Asparagic acid | HMDB | | Asparaginate | HMDB | | Asparaginic acid | HMDB | | Asparatate | HMDB | | H-Asp-OH | HMDB | | L-(+)-Aspartate | HMDB | | L-(+)-Aspartic acid | HMDB | | L-Aminosuccinate | HMDB | | L-Aminosuccinic acid | HMDB | | L-Asparagate | HMDB | | L-Asparagic acid | HMDB | | L-Asparaginate | HMDB | | L-Asparaginic acid | HMDB | | (+-)-Aspartic acid | HMDB | | (R,S)-Aspartic acid | HMDB | | Aspartate, disodium | HMDB | | Aspartate, magnesium | HMDB | | Aspartate, monopotassium | HMDB | | Aspartic acid, dipotassium salt | HMDB | | Aspartic acid, hydrobromide | HMDB | | Aspartic acid, monopotassium salt | HMDB | | Aspartic acid, monosodium salt | HMDB | | Aspartic acid, potassium salt | HMDB | | L Aspartate | HMDB | | MG5Longoral | HMDB | | Potassium aspartate | HMDB | | Polysuccinimide | HMDB | | Ammonium aspartate | HMDB | | Aspartate, ammonium | HMDB | | Aspartate, calcium | HMDB | | Aspartate, monosodium | HMDB | | Aspartic acid, calcium salt | HMDB | | Aspartic acid, disodium salt | HMDB | | Aspartic acid, magnesium (1:1) salt, hydrochloride, trihydrate | HMDB | | Dipotassium aspartate | HMDB | | Disodium aspartate | HMDB | | Hydrochloride, aspartate magnesium | HMDB | | Monopotassium aspartate | HMDB | | Sodium aspartate | HMDB | | Aspartate, dipotassium | HMDB | | Aspartic acid, magnesium (2:1) salt | HMDB | | Aspartic acid, sodium salt | HMDB | | Hydrobromide aspartic acid | HMDB | | Magnesium aspartate | HMDB | | MG 5 Longoral | HMDB | | Monosodium aspartate | HMDB | | Aspartate magnesium hydrochloride | HMDB | | Aspartate, potassium | HMDB | | Aspartate, sodium | HMDB | | Aspartic acid, ammonium salt | HMDB | | Aspartic acid, hydrochloride | HMDB | | Aspartic acid, magnesium-potassium (2:1:2) salt | HMDB | | Calcium aspartate | HMDB | | Hydrochloride aspartic acid | HMDB | | L Aspartic acid | HMDB | | Magnesiocard | HMDB | | MG-5-Longoral | HMDB | | Poly-DL-succinimide | HMDB |
| Show more...
|---|
| Chemical Formula | C4H7NO4 |
|---|
| Average Molecular Weight | 133.1027 |
|---|
| Monoisotopic Molecular Weight | 133.037507717 |
|---|
| IUPAC Name | (2S)-2-aminobutanedioic acid |
|---|
| Traditional Name | L-aspartic acid |
|---|
| CAS Registry Number | 56-84-8 |
|---|
| SMILES | N[C@@H](CC(O)=O)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C4H7NO4/c5-2(4(8)9)1-3(6)7/h2H,1,5H2,(H,6,7)(H,8,9)/t2-/m0/s1 |
|---|
| InChI Key | CKLJMWTZIZZHCS-REOHCLBHSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as aspartic acid and derivatives. Aspartic acid and derivatives are compounds containing an aspartic acid or a derivative thereof resulting from reaction of aspartic acid at the amino group or the carboxy group, or from the replacement of any hydrogen of glycine by a heteroatom. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Carboxylic acids and derivatives |
|---|
| Sub Class | Amino acids, peptides, and analogues |
|---|
| Direct Parent | Aspartic acid and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Aspartic acid or derivatives
- Alpha-amino acid
- L-alpha-amino acid
- Dicarboxylic acid or derivatives
- Fatty acid
- Amino acid
- Carboxylic acid
- Organic oxide
- Organopnictogen compound
- Primary amine
- Organooxygen compound
- Organonitrogen compound
- Primary aliphatic amine
- Organic oxygen compound
- Carbonyl group
- Amine
- Organic nitrogen compound
- Hydrocarbon derivative
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Physiological effect | |
|---|
| Disposition | Biological locationRoute of exposureSourceEndogenousExogenousFood- Food (HMDB: HMDB0000191)
Animal originHerb and spiceVegetableFruitNutCereal and cereal productPulseGourdCoffee and coffee productSoyDishTeaBaking goodBeverageAquatic originBaby foodUnclassified food or beverageFat and oilMilk and milk productOther milk productFermented milk productUnfermented milk- Milk (Other mammals) (FooDB: FOOD00690)
- Milk (Human) (FooDB: FOOD00666)
- Milk (Cow) (FooDB: FOOD00618)
- Cow milk, pasteurized, vitamin A + D added, 0% fat (FooDB: FOOD00889)
- Cow milk, pasteurized, vitamin A + D added, 1% fat (FooDB: FOOD00890)
- Cow milk, pasteurized, vitamin A + D added, 2% fat (FooDB: FOOD00891)
- Cow milk, pasteurized, vitamin D added, 3.25% fat (FooDB: FOOD00892)
Fermented milk Cocoa and cocoa productSnackConfectioneryEgg
|
|---|
| Process | |
|---|
| Role | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | 270 °C | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | 5.39 mg/mL | YALKOWSKY,SH & DANNENFELSER,RM (1992) | | LogP | -3.89 | CHMELIK,J ET AL. (1991) |
|
|---|
| Experimental Chromatographic Properties | Experimental Collision Cross Sections |
|---|
| Predicted Molecular Properties | | Show more...
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Kovats Retention IndicesUnderivatizedDerivatized| Derivative Name / Structure | SMILES | Kovats RI Value | Column Type | Reference |
|---|
| L-Aspartic acid,1TMS,isomer #1 | C[Si](C)(C)OC(=O)C[C@H](N)C(=O)O | 1393.4 | Semi standard non polar | 33892256 | | L-Aspartic acid,1TMS,isomer #2 | C[Si](C)(C)OC(=O)[C@@H](N)CC(=O)O | 1347.9 | Semi standard non polar | 33892256 | | L-Aspartic acid,1TMS,isomer #3 | C[Si](C)(C)N[C@@H](CC(=O)O)C(=O)O | 1412.7 | Semi standard non polar | 33892256 | | L-Aspartic acid,2TMS,isomer #1 | C[Si](C)(C)OC(=O)C[C@H](N)C(=O)O[Si](C)(C)C | 1413.6 | Semi standard non polar | 33892256 | | L-Aspartic acid,2TMS,isomer #2 | C[Si](C)(C)N[C@@H](CC(=O)O[Si](C)(C)C)C(=O)O | 1504.6 | Semi standard non polar | 33892256 | | L-Aspartic acid,2TMS,isomer #3 | C[Si](C)(C)N[C@@H](CC(=O)O)C(=O)O[Si](C)(C)C | 1457.1 | Semi standard non polar | 33892256 | | L-Aspartic acid,2TMS,isomer #4 | C[Si](C)(C)N([C@@H](CC(=O)O)C(=O)O)[Si](C)(C)C | 1619.8 | Semi standard non polar | 33892256 | | L-Aspartic acid,3TMS,isomer #1 | C[Si](C)(C)N[C@@H](CC(=O)O[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1548.8 | Semi standard non polar | 33892256 | | L-Aspartic acid,3TMS,isomer #1 | C[Si](C)(C)N[C@@H](CC(=O)O[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1566.3 | Standard non polar | 33892256 | | L-Aspartic acid,3TMS,isomer #1 | C[Si](C)(C)N[C@@H](CC(=O)O[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1715.5 | Standard polar | 33892256 | | L-Aspartic acid,3TMS,isomer #2 | C[Si](C)(C)OC(=O)C[C@@H](C(=O)O)N([Si](C)(C)C)[Si](C)(C)C | 1651.1 | Semi standard non polar | 33892256 | | L-Aspartic acid,3TMS,isomer #2 | C[Si](C)(C)OC(=O)C[C@@H](C(=O)O)N([Si](C)(C)C)[Si](C)(C)C | 1611.5 | Standard non polar | 33892256 | | L-Aspartic acid,3TMS,isomer #2 | C[Si](C)(C)OC(=O)C[C@@H](C(=O)O)N([Si](C)(C)C)[Si](C)(C)C | 1863.4 | Standard polar | 33892256 | | L-Aspartic acid,3TMS,isomer #3 | C[Si](C)(C)OC(=O)[C@H](CC(=O)O)N([Si](C)(C)C)[Si](C)(C)C | 1628.8 | Semi standard non polar | 33892256 | | L-Aspartic acid,3TMS,isomer #3 | C[Si](C)(C)OC(=O)[C@H](CC(=O)O)N([Si](C)(C)C)[Si](C)(C)C | 1614.9 | Standard non polar | 33892256 | | L-Aspartic acid,3TMS,isomer #3 | C[Si](C)(C)OC(=O)[C@H](CC(=O)O)N([Si](C)(C)C)[Si](C)(C)C | 1811.5 | Standard polar | 33892256 | | L-Aspartic acid,4TMS,isomer #1 | C[Si](C)(C)OC(=O)C[C@@H](C(=O)O[Si](C)(C)C)N([Si](C)(C)C)[Si](C)(C)C | 1707.3 | Semi standard non polar | 33892256 | | L-Aspartic acid,4TMS,isomer #1 | C[Si](C)(C)OC(=O)C[C@@H](C(=O)O[Si](C)(C)C)N([Si](C)(C)C)[Si](C)(C)C | 1680.4 | Standard non polar | 33892256 | | L-Aspartic acid,4TMS,isomer #1 | C[Si](C)(C)OC(=O)C[C@@H](C(=O)O[Si](C)(C)C)N([Si](C)(C)C)[Si](C)(C)C | 1664.0 | Standard polar | 33892256 | | L-Aspartic acid,1TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)C[C@H](N)C(=O)O | 1626.3 | Semi standard non polar | 33892256 | | L-Aspartic acid,1TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC(=O)[C@@H](N)CC(=O)O | 1598.8 | Semi standard non polar | 33892256 | | L-Aspartic acid,1TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)N[C@@H](CC(=O)O)C(=O)O | 1672.0 | Semi standard non polar | 33892256 | | L-Aspartic acid,2TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)C[C@H](N)C(=O)O[Si](C)(C)C(C)(C)C | 1857.9 | Semi standard non polar | 33892256 | | L-Aspartic acid,2TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)N[C@@H](CC(=O)O[Si](C)(C)C(C)(C)C)C(=O)O | 1937.4 | Semi standard non polar | 33892256 | | L-Aspartic acid,2TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)N[C@@H](CC(=O)O)C(=O)O[Si](C)(C)C(C)(C)C | 1898.5 | Semi standard non polar | 33892256 | | L-Aspartic acid,2TBDMS,isomer #4 | CC(C)(C)[Si](C)(C)N([C@@H](CC(=O)O)C(=O)O)[Si](C)(C)C(C)(C)C | 2057.2 | Semi standard non polar | 33892256 | | L-Aspartic acid,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)N[C@@H](CC(=O)O[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2153.5 | Semi standard non polar | 33892256 | | L-Aspartic acid,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)N[C@@H](CC(=O)O[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2162.8 | Standard non polar | 33892256 | | L-Aspartic acid,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)N[C@@H](CC(=O)O[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2113.0 | Standard polar | 33892256 | | L-Aspartic acid,3TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC(=O)C[C@@H](C(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2292.4 | Semi standard non polar | 33892256 | | L-Aspartic acid,3TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC(=O)C[C@@H](C(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2222.1 | Standard non polar | 33892256 | | L-Aspartic acid,3TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC(=O)C[C@@H](C(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2173.3 | Standard polar | 33892256 | | L-Aspartic acid,3TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CC(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2301.2 | Semi standard non polar | 33892256 | | L-Aspartic acid,3TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CC(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2201.8 | Standard non polar | 33892256 | | L-Aspartic acid,3TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CC(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2143.9 | Standard polar | 33892256 | | L-Aspartic acid,4TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)C[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2544.7 | Semi standard non polar | 33892256 | | L-Aspartic acid,4TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)C[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2441.4 | Standard non polar | 33892256 | | L-Aspartic acid,4TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)C[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2169.3 | Standard polar | 33892256 |
| Show more...
|---|
| Spectra |
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-0f89-0960000000-5a6a6cb21e9fc0875f84 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-0f89-0950000000-51399a8446c394459765 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-0f89-0950000000-f07e7f52b3c31fd119f5 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-0f89-0950000000-9e175dfa17a8b17a72d2 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (Non-derivatized) | splash10-0f89-0950000000-f4475587d5a3b20bfc58 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-00di-9430000000-0cac9a97e698db1672b5 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-MS (2 TMS) | splash10-03yi-1900000000-ea61b8526ee02ae79b66 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-MS (3 TMS) | splash10-0f89-1890000000-7da576c8129142b71a1b | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-MS (4 TMS) | splash10-0pb9-0981000000-ad23d55e348f55115f00 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid EI-B (Non-derivatized) | splash10-001i-0590000000-13cc5a7841854e97f68b | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-EI-TOF (Non-derivatized) | splash10-0f89-0960000000-5a6a6cb21e9fc0875f84 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-EI-TOF (Non-derivatized) | splash10-0f89-0950000000-51399a8446c394459765 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-EI-TOF (Non-derivatized) | splash10-0f89-0950000000-f07e7f52b3c31fd119f5 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-EI-TOF (Non-derivatized) | splash10-0f89-0950000000-9e175dfa17a8b17a72d2 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-EI-TOF (Non-derivatized) | splash10-0f89-0950000000-f4475587d5a3b20bfc58 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-EI-QQ (Non-derivatized) | splash10-0g4i-6946100000-0ae49e3be0f2f1f92065 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-EI-TOF (Non-derivatized) | splash10-00di-9430000000-0cac9a97e698db1672b5 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-MS (Non-derivatized) | splash10-03yi-1900000000-ea61b8526ee02ae79b66 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-MS (Non-derivatized) | splash10-0f89-1890000000-7da576c8129142b71a1b | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - L-Aspartic acid GC-MS (Non-derivatized) | splash10-0pb9-0981000000-ad23d55e348f55115f00 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - L-Aspartic acid GC-MS (Non-derivatized) - 70eV, Positive | splash10-000f-9100000000-b7db3377089f4c95001b | 2016-09-22 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - L-Aspartic acid GC-MS (2 TMS) - 70eV, Positive | splash10-01w4-9610000000-0f1463c6395830708e12 | 2017-10-06 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - L-Aspartic acid GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - L-Aspartic acid GC-MS (TMS_1_1) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - L-Aspartic acid GC-MS (TMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid Quattro_QQQ 10V, Positive-QTOF (Annotated) | splash10-0079-9300000000-92f9914d94078d96f5cf | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid Quattro_QQQ 25V, Positive-QTOF (Annotated) | splash10-00di-9000000000-c467d10f2b7e21ed1734 | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid Quattro_QQQ 40V, Positive-QTOF (Annotated) | splash10-00dl-9000000000-87f0c58226f5a2f3ac8a | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-00di-0900000000-fde7ef1951fddff4b817 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-0uk9-8900000000-2d7e5609618437e59272 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-000i-9000000000-4b43567f4a446aed0828 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-014i-4900000000-45382d9abd25be948e5b | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-001i-0900000000-3f39a9b758e282358ac0 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-014i-3900000000-44d361ad09ff9a30dd14 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-000i-9000000000-18e1dba62e6b803e17b8 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-0002-0930000000-d97f9518a2d516d830c5 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-02ai-0962100000-909bb894b2c1318afee4 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-00di-9000000000-f2059dd438fcf62f62e1 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-03dr-7900000000-8b7b89ed34530e331024 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-014i-0190000000-58a78949f4b93de22aab | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-001i-0921000000-214b6b969fdc4216ca71 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-03dr-6900000000-4918748927dd097d2879 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-001i-0900000000-88dc2f1093f261e76201 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-004i-0290000000-a8856f56645a2961baac | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-QQ (API3000, Applied Biosystems) 10V, Negative-QTOF | splash10-001i-0900000000-8cf9870557ca9adc3374 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-QQ (API3000, Applied Biosystems) 20V, Negative-QTOF | splash10-000i-9400000000-81b2804a712625a2d9c4 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-QQ (API3000, Applied Biosystems) 30V, Negative-QTOF | splash10-000i-9000000000-9bf77ba12ad952f84ac0 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-QQ (API3000, Applied Biosystems) 40V, Negative-QTOF | splash10-00di-9000000000-e08bfb96c5fd8c9b9db7 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-QQ (API3000, Applied Biosystems) 50V, Negative-QTOF | splash10-0006-9000000000-876e30f6c9ed061091fe | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - L-Aspartic acid LC-ESI-QQ (API3000, Applied Biosystems) 10V, Positive-QTOF | splash10-00dl-9000000000-6c0d44d4e3853e5701a3 | 2012-08-31 | HMDB team, MONA | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Experimental 1D NMR | 1H NMR Spectrum (1D, 600 MHz, H2O, experimental) | 2012-12-04 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-16 | Wishart Lab | View Spectrum | | Experimental 1D NMR | 13C NMR Spectrum (1D, 400 MHz, H2O, experimental) | 2021-10-10 | Wishart Lab | View Spectrum | | Experimental 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 400 MHz, H2O, experimental) | 2012-12-05 | Wishart Lab | View Spectrum |
IR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M-H]-) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+H]+) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+Na]+) | 2023-02-03 | FELIX lab | View Spectrum |
| Show more...
|---|
| Biological Properties |
|---|
| Cellular Locations | - Cytoplasm
- Extracellular
- Mitochondria
|
|---|
| Biospecimen Locations | - Blood
- Breast Milk
- Cerebrospinal Fluid (CSF)
- Feces
- Saliva
- Sweat
- Urine
|
|---|
| Tissue Locations | - All Tissues
- Placenta
- Prostate
|
|---|
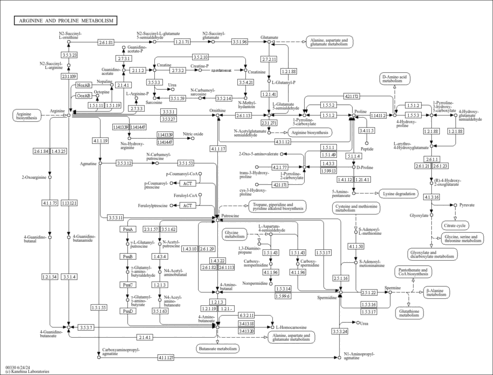
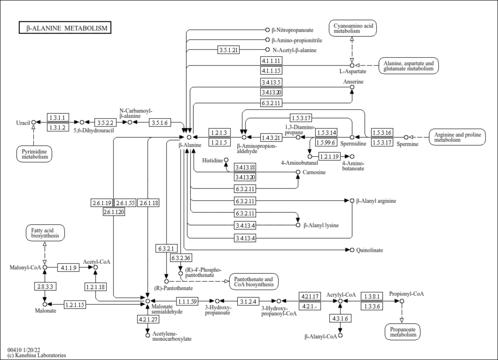
| Pathways | |
|---|
| Normal Concentrations |
|---|
| |
| Blood | Detected and Quantified | 21.0 +/- 5.0 uM | Adult (>18 years old) | Male | Normal | | details | | Blood | Detected and Quantified | 20.0 +/- 5.0 uM | Adult (>18 years old) | Female | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 15.5 (tr-32.1) uM | Infant (0-1 year old) | Not Specified | Normal | | details | | Blood | Detected and Quantified | 6.26 +/- 2.34 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 8.00-20.0 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 54.4(47.3-68.3) uM | Children (1-13 uears old) | Both | Normal | | details | | Blood | Detected and Quantified | 20.9 +/- 6.1 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Breast Milk | Detected and Quantified | 52.9 +/- 33.6 uM | Adult (>18 years old) | Female | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 2.30 (0.0-4.6) uM | Adult (>18 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 2.8 +/- 1.2 uM | Adult (>18 years old) | Male | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 1.8 +/- 0.2 uM | Adult (>18 years old) | Female | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 0.6 +/- 0.3 uM | Adult (>18 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 0.6 +/- 0.3 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 0.9 +/- 0.5 uM | Infant (0-1 year old) | Not Specified | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 2.19 +/- 2.63 uM | Adult (>18 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 0.23 +/- 0.18 uM | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Not Specified | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected and Quantified | 90 +/- 90 nmol/g wet feces | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected and Quantified | 130 +/- 120 nmol/g wet feces | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Not Specified | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected and Quantified | 6.54 +/- 2.35 uM | Adult (>18 years old) | Female | Normal | | details | | Saliva | Detected and Quantified | 18.2 +/- 14.6 uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 2.6 +/- 3.6 uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 1.5 +/- 2.1 uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | <0.25 uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | >10 uM | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected and Quantified | 33.30 +/- 19.39 uM | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected and Quantified | <0.25 uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 13.83 +/- 5.06 uM | Adult (>18 years old) | Female | Normal | | details | | Sweat | Detected and Quantified | < 10 uM | Adult (60 years old) | Male | Normal | | details | | Sweat | Detected and Quantified | < 10 uM | Adult (40 years old) | Male | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 5.57-34.61 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 16.85 +/- 7.16 umol/mmol creatinine | Newborn (0-30 days old) | Female | Normal | | details | | Urine | Detected and Quantified | 16.88 +/- 7.25 umol/mmol creatinine | Newborn (0-30 days old) | Male | Normal | | details | | Urine | Detected and Quantified | 0.43-0.82 umol/mmol creatinine | Adult (>18 years old) | Female | Normal | | details | | Urine | Detected and Quantified | 1.36 +/- 1.06 umol/mmol creatinine | Infant (0-1 year old) | Both | Normal | | details | | Urine | Detected and Quantified | 25.11 (20-30.22) umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | - Geigy Scientific ...
- West Cadwell, N.J...
- Basel, Switzerlan...
| details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 10.9 (3.5-21.8) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 16.115 +/- 7.486 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 5.2 +/- 6.15 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 0.4 +/- 0.2 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 0.44-0.68 umol/mmol creatinine | Adult (>18 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 0.5 +/-0.4 umol/mmol creatinine | Infant (1 - 6 months old) | Both | Normal | | details | | Urine | Detected and Quantified | 0.6 +/- 0.3 umol/mmol creatinine | Infant (6 months - <1 year old) | Both | Normal | | details | | Urine | Detected and Quantified | 10.9 (1.9-26.8) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 14.27 (9.19-21.64) umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 0.5 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 51.1451 +/- 42.3486 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 4.638 (1.711-7.566) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 16.62 +/- 12.44 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 18.20 +/- 13.57 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 18.20 +/- 18.09 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details |
| Show more...
|---|
| Abnormal Concentrations |
|---|
| |
| Blood | Detected and Quantified | 36.0 +/- 9.0 uM | Adult (>18 years old) | Both | Cirrhosis | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Blood | Detected and Quantified | 3.9 (3.7-4.1) uM | Adult (>18 years old) | Both | Epilepsy | | details | | Blood | Detected and Quantified | 11.8 (10.9-12.7) uM | Adult (>18 years old) | Both | Epilepsy | | details | | Blood | Detected and Quantified | 2.6 (tr-5.0) uM | Infant (0-1 year old) | Male | Dicarboxylic Aminoaciduria | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Blood | Detected and Quantified | 56.7(42.3-70.3) uM | Children (1-13 uears old) | Both | Environmental enteric dysfunction | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Schizophrenia | | details | | Blood | Detected and Quantified | 12.67 +/- 2.86 uM | Elderly (>65 years old) | Both | Alzheimer's disease | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Schizophrenia | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 0.233 +/- 0.081 uM | Adult (>18 years old) | Not Specified | Growth hormone deficiency | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | trace uM | Infant (0-1 year old) | Male | Dicarboxylic Aminoaciduria | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 0.24 +/- 0.10 uM | Adult (>18 years old) | Both | Schizophrenia | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Gout | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal Cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Irritable bowel syndrome | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Ulcerative colitis | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Immunoglobulin A nephropathy (IgAN) progressor | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Not Specified | Not Specified | Cryptosporidium infection | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Both | Crohns disease | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Both | Ulcerative colitis | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Not Specified | Crohns disease | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Not Specified | Ulcerative colitis | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Not Specified | Unclassified IBD | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Crohns disease | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Ulcerative colitis | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Attachment loss | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Missing teeth | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Periodontal Probing Depth | | details | | Saliva | Detected and Quantified | 9.62 +/- 6.59 uM | Adult (>18 years old) | Male | Alzheimer's disease | | details | | Saliva | Detected and Quantified | 11.32 +/- 3.39 uM | Adult (>18 years old) | Male | Frontotemporal lobe dementia | | details | | Saliva | Detected and Quantified | 13.36 +/- 3.74 uM | Adult (>18 years old) | Both | Lewy body disease | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Oral cancer | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | Breast cancer | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Pancreatic cancer | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Periodontal diseases | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Bladder cancer | | details | | Urine | Detected and Quantified | 24.087 +/- 22.064 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Eosinophilic esophagitis | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Urine | Detected and Quantified | 29.311 +/- 17.1956 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Eosinophilic esophagitis | | details | | Urine | Detected and Quantified | 28.1856 +/- 25.7291 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Gastroesophageal reflux disease | | details |
| Show more...
|---|
| Associated Disorders and Diseases |
|---|
| Disease References | | Epilepsy |
|---|
- Rainesalo S, Keranen T, Palmio J, Peltola J, Oja SS, Saransaari P: Plasma and cerebrospinal fluid amino acids in epileptic patients. Neurochem Res. 2004 Jan;29(1):319-24. [PubMed:14992292 ]
| | Cirrhosis |
|---|
- Cynober LA: Plasma amino acid levels with a note on membrane transport: characteristics, regulation, and metabolic significance. Nutrition. 2002 Sep;18(9):761-6. [PubMed:12297216 ]
| | Alzheimer's disease |
|---|
- Fonteh AN, Harrington RJ, Tsai A, Liao P, Harrington MG: Free amino acid and dipeptide changes in the body fluids from Alzheimer's disease subjects. Amino Acids. 2007 Feb;32(2):213-24. Epub 2006 Oct 10. [PubMed:17031479 ]
- Tsuruoka M, Hara J, Hirayama A, Sugimoto M, Soga T, Shankle WR, Tomita M: Capillary electrophoresis-mass spectrometry-based metabolome analysis of serum and saliva from neurodegenerative dementia patients. Electrophoresis. 2013 Oct;34(19):2865-72. doi: 10.1002/elps.201300019. Epub 2013 Sep 6. [PubMed:23857558 ]
| | Dicarboxylic aminoaciduria |
|---|
- Melancon SB, Dallaire L, Lemieux B, Robitaille P, Potier M: Dicarboxylic aminoaciduria: an inborn error of amino acid conservation. J Pediatr. 1977 Sep;91(3):422-7. [PubMed:894411 ]
| | Growth hormone deficiency |
|---|
- Burman P, Hetta J, Wide L, Mansson JE, Ekman R, Karlsson FA: Growth hormone treatment affects brain neurotransmitters and thyroxine [see comment]. Clin Endocrinol (Oxf). 1996 Mar;44(3):319-24. [PubMed:8729530 ]
| | Schizophrenia |
|---|
- Do KQ, Lauer CJ, Schreiber W, Zollinger M, Gutteck-Amsler U, Cuenod M, Holsboer F: gamma-Glutamylglutamine and taurine concentrations are decreased in the cerebrospinal fluid of drug-naive patients with schizophrenic disorders. J Neurochem. 1995 Dec;65(6):2652-62. [PubMed:7595563 ]
- Xuan J, Pan G, Qiu Y, Yang L, Su M, Liu Y, Chen J, Feng G, Fang Y, Jia W, Xing Q, He L: Metabolomic profiling to identify potential serum biomarkers for schizophrenia and risperidone action. J Proteome Res. 2011 Dec 2;10(12):5433-43. doi: 10.1021/pr2006796. Epub 2011 Nov 8. [PubMed:22007635 ]
- Yang J, Chen T, Sun L, Zhao Z, Qi X, Zhou K, Cao Y, Wang X, Qiu Y, Su M, Zhao A, Wang P, Yang P, Wu J, Feng G, He L, Jia W, Wan C: Potential metabolite markers of schizophrenia. Mol Psychiatry. 2013 Jan;18(1):67-78. doi: 10.1038/mp.2011.131. Epub 2011 Oct 25. [PubMed:22024767 ]
| | Irritable bowel syndrome |
|---|
- Le Gall G, Noor SO, Ridgway K, Scovell L, Jamieson C, Johnson IT, Colquhoun IJ, Kemsley EK, Narbad A: Metabolomics of fecal extracts detects altered metabolic activity of gut microbiota in ulcerative colitis and irritable bowel syndrome. J Proteome Res. 2011 Sep 2;10(9):4208-18. doi: 10.1021/pr2003598. Epub 2011 Aug 8. [PubMed:21761941 ]
| | Ulcerative colitis |
|---|
- Le Gall G, Noor SO, Ridgway K, Scovell L, Jamieson C, Johnson IT, Colquhoun IJ, Kemsley EK, Narbad A: Metabolomics of fecal extracts detects altered metabolic activity of gut microbiota in ulcerative colitis and irritable bowel syndrome. J Proteome Res. 2011 Sep 2;10(9):4208-18. doi: 10.1021/pr2003598. Epub 2011 Aug 8. [PubMed:21761941 ]
- Bjerrum JT, Wang Y, Hao F, Coskun M, Ludwig C, Gunther U, Nielsen OH: Metabonomics of human fecal extracts characterize ulcerative colitis, Crohn's disease and healthy individuals. Metabolomics. 2015;11:122-133. Epub 2014 Jun 1. [PubMed:25598765 ]
- Kolho KL, Pessia A, Jaakkola T, de Vos WM, Velagapudi V: Faecal and Serum Metabolomics in Paediatric Inflammatory Bowel Disease. J Crohns Colitis. 2017 Mar 1;11(3):321-334. doi: 10.1093/ecco-jcc/jjw158. [PubMed:27609529 ]
| | Colorectal cancer |
|---|
- Weir TL, Manter DK, Sheflin AM, Barnett BA, Heuberger AL, Ryan EP: Stool microbiome and metabolome differences between colorectal cancer patients and healthy adults. PLoS One. 2013 Aug 6;8(8):e70803. doi: 10.1371/journal.pone.0070803. Print 2013. [PubMed:23940645 ]
- Ritchie SA, Ahiahonu PW, Jayasinghe D, Heath D, Liu J, Lu Y, Jin W, Kavianpour A, Yamazaki Y, Khan AM, Hossain M, Su-Myat KK, Wood PL, Krenitsky K, Takemasa I, Miyake M, Sekimoto M, Monden M, Matsubara H, Nomura F, Goodenowe DB: Reduced levels of hydroxylated, polyunsaturated ultra long-chain fatty acids in the serum of colorectal cancer patients: implications for early screening and detection. BMC Med. 2010 Feb 15;8:13. doi: 10.1186/1741-7015-8-13. [PubMed:20156336 ]
- Ni Y, Xie G, Jia W: Metabonomics of human colorectal cancer: new approaches for early diagnosis and biomarker discovery. J Proteome Res. 2014 Sep 5;13(9):3857-70. doi: 10.1021/pr500443c. Epub 2014 Aug 14. [PubMed:25105552 ]
- Brown DG, Rao S, Weir TL, O'Malia J, Bazan M, Brown RJ, Ryan EP: Metabolomics and metabolic pathway networks from human colorectal cancers, adjacent mucosa, and stool. Cancer Metab. 2016 Jun 6;4:11. doi: 10.1186/s40170-016-0151-y. eCollection 2016. [PubMed:27275383 ]
- Sinha R, Ahn J, Sampson JN, Shi J, Yu G, Xiong X, Hayes RB, Goedert JJ: Fecal Microbiota, Fecal Metabolome, and Colorectal Cancer Interrelations. PLoS One. 2016 Mar 25;11(3):e0152126. doi: 10.1371/journal.pone.0152126. eCollection 2016. [PubMed:27015276 ]
- Goedert JJ, Sampson JN, Moore SC, Xiao Q, Xiong X, Hayes RB, Ahn J, Shi J, Sinha R: Fecal metabolomics: assay performance and association with colorectal cancer. Carcinogenesis. 2014 Sep;35(9):2089-96. doi: 10.1093/carcin/bgu131. Epub 2014 Jul 18. [PubMed:25037050 ]
- Wang X, Wang J, Rao B, Deng L: Gut flora profiling and fecal metabolite composition of colorectal cancer patients and healthy individuals. Exp Ther Med. 2017 Jun;13(6):2848-2854. doi: 10.3892/etm.2017.4367. Epub 2017 Apr 20. [PubMed:28587349 ]
| | Crohn's disease |
|---|
- Bjerrum JT, Wang Y, Hao F, Coskun M, Ludwig C, Gunther U, Nielsen OH: Metabonomics of human fecal extracts characterize ulcerative colitis, Crohn's disease and healthy individuals. Metabolomics. 2015;11:122-133. Epub 2014 Jun 1. [PubMed:25598765 ]
- Kolho KL, Pessia A, Jaakkola T, de Vos WM, Velagapudi V: Faecal and Serum Metabolomics in Paediatric Inflammatory Bowel Disease. J Crohns Colitis. 2017 Mar 1;11(3):321-334. doi: 10.1093/ecco-jcc/jjw158. [PubMed:27609529 ]
| | Gout |
|---|
- Shao T, Shao L, Li H, Xie Z, He Z, Wen C: Combined Signature of the Fecal Microbiome and Metabolome in Patients with Gout. Front Microbiol. 2017 Feb 21;8:268. doi: 10.3389/fmicb.2017.00268. eCollection 2017. [PubMed:28270806 ]
| | Perillyl alcohol administration for cancer treatment |
|---|
- Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M: Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics. 2010 Mar;6(1):78-95. Epub 2009 Sep 10. [PubMed:20300169 ]
| | Pancreatic cancer |
|---|
- Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M: Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics. 2010 Mar;6(1):78-95. Epub 2009 Sep 10. [PubMed:20300169 ]
| | Periodontal disease |
|---|
- Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M: Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics. 2010 Mar;6(1):78-95. Epub 2009 Sep 10. [PubMed:20300169 ]
| | Frontotemporal dementia |
|---|
- Tsuruoka M, Hara J, Hirayama A, Sugimoto M, Soga T, Shankle WR, Tomita M: Capillary electrophoresis-mass spectrometry-based metabolome analysis of serum and saliva from neurodegenerative dementia patients. Electrophoresis. 2013 Oct;34(19):2865-72. doi: 10.1002/elps.201300019. Epub 2013 Sep 6. [PubMed:23857558 ]
| | Lewy body disease |
|---|
- Tsuruoka M, Hara J, Hirayama A, Sugimoto M, Soga T, Shankle WR, Tomita M: Capillary electrophoresis-mass spectrometry-based metabolome analysis of serum and saliva from neurodegenerative dementia patients. Electrophoresis. 2013 Oct;34(19):2865-72. doi: 10.1002/elps.201300019. Epub 2013 Sep 6. [PubMed:23857558 ]
| | Attachment loss |
|---|
- Liebsch C, Pitchika V, Pink C, Samietz S, Kastenmuller G, Artati A, Suhre K, Adamski J, Nauck M, Volzke H, Friedrich N, Kocher T, Holtfreter B, Pietzner M: The Saliva Metabolome in Association to Oral Health Status. J Dent Res. 2019 Jun;98(6):642-651. doi: 10.1177/0022034519842853. Epub 2019 Apr 26. [PubMed:31026179 ]
| | Missing teeth |
|---|
- Liebsch C, Pitchika V, Pink C, Samietz S, Kastenmuller G, Artati A, Suhre K, Adamski J, Nauck M, Volzke H, Friedrich N, Kocher T, Holtfreter B, Pietzner M: The Saliva Metabolome in Association to Oral Health Status. J Dent Res. 2019 Jun;98(6):642-651. doi: 10.1177/0022034519842853. Epub 2019 Apr 26. [PubMed:31026179 ]
| | Periodontal Probing Depth |
|---|
- Liebsch C, Pitchika V, Pink C, Samietz S, Kastenmuller G, Artati A, Suhre K, Adamski J, Nauck M, Volzke H, Friedrich N, Kocher T, Holtfreter B, Pietzner M: The Saliva Metabolome in Association to Oral Health Status. J Dent Res. 2019 Jun;98(6):642-651. doi: 10.1177/0022034519842853. Epub 2019 Apr 26. [PubMed:31026179 ]
| | Eosinophilic esophagitis |
|---|
- Slae, M., Huynh, H., Wishart, D.S. (2014). Analysis of 30 normal pediatric urine samples via NMR spectroscopy (unpublished work). NA.
|
|
|---|
| Associated OMIM IDs | |
|---|
| External Links |
|---|
| DrugBank ID | DB00128 |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FooDB ID | FDB012567 |
|---|
| KNApSAcK ID | C00001342 |
|---|
| Chemspider ID | 5745 |
|---|
| KEGG Compound ID | C00049 |
|---|
| BioCyc ID | L-ASPARTATE |
|---|
| BiGG ID | 33663 |
|---|
| Wikipedia Link | Aspartic acid |
|---|
| METLIN ID | 5206 |
|---|
| PubChem Compound | 5960 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 17053 |
|---|
| Food Biomarker Ontology | Not Available |
|---|
| VMH ID | ASP_L |
|---|
| MarkerDB ID | MDB00000091 |
|---|
| Good Scents ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Pamfil, Maria; Lupescu, Irina; Savoiu, Valeria Gabriela. L-aspartic acid production from fumarate using Escherichia coli whole cells. Rom. (2005), 3pp. |
|---|
| Material Safety Data Sheet (MSDS) | Not Available |
|---|
| General References | - Sreekumar A, Poisson LM, Rajendiran TM, Khan AP, Cao Q, Yu J, Laxman B, Mehra R, Lonigro RJ, Li Y, Nyati MK, Ahsan A, Kalyana-Sundaram S, Han B, Cao X, Byun J, Omenn GS, Ghosh D, Pennathur S, Alexander DC, Berger A, Shuster JR, Wei JT, Varambally S, Beecher C, Chinnaiyan AM: Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature. 2009 Feb 12;457(7231):910-4. doi: 10.1038/nature07762. [PubMed:19212411 ]
- Silwood CJ, Lynch E, Claxson AW, Grootveld MC: 1H and (13)C NMR spectroscopic analysis of human saliva. J Dent Res. 2002 Jun;81(6):422-7. [PubMed:12097436 ]
- Nicholson JK, O'Flynn MP, Sadler PJ, Macleod AF, Juul SM, Sonksen PH: Proton-nuclear-magnetic-resonance studies of serum, plasma and urine from fasting normal and diabetic subjects. Biochem J. 1984 Jan 15;217(2):365-75. [PubMed:6696735 ]
- Engelborghs S, Marescau B, De Deyn PP: Amino acids and biogenic amines in cerebrospinal fluid of patients with Parkinson's disease. Neurochem Res. 2003 Aug;28(8):1145-50. [PubMed:12834252 ]
- Burman P, Hetta J, Wide L, Mansson JE, Ekman R, Karlsson FA: Growth hormone treatment affects brain neurotransmitters and thyroxine [see comment]. Clin Endocrinol (Oxf). 1996 Mar;44(3):319-24. [PubMed:8729530 ]
- Hagenfeldt L, Bjerkenstedt L, Edman G, Sedvall G, Wiesel FA: Amino acids in plasma and CSF and monoamine metabolites in CSF: interrelationship in healthy subjects. J Neurochem. 1984 Mar;42(3):833-7. [PubMed:6198473 ]
- Cynober LA: Plasma amino acid levels with a note on membrane transport: characteristics, regulation, and metabolic significance. Nutrition. 2002 Sep;18(9):761-6. [PubMed:12297216 ]
- Rainesalo S, Keranen T, Palmio J, Peltola J, Oja SS, Saransaari P: Plasma and cerebrospinal fluid amino acids in epileptic patients. Neurochem Res. 2004 Jan;29(1):319-24. [PubMed:14992292 ]
- Chiara F, Goumans MJ, Forsberg H, Ahgren A, Rasola A, Aspenstrom P, Wernstedt C, Hellberg C, Heldin CH, Heuchel R: A gain of function mutation in the activation loop of platelet-derived growth factor beta-receptor deregulates its kinase activity. J Biol Chem. 2004 Oct 8;279(41):42516-27. Epub 2004 Jul 28. [PubMed:15284236 ]
- Fujii N: D-amino acid in elderly tissues. Biol Pharm Bull. 2005 Sep;28(9):1585-9. [PubMed:16141520 ]
- Grdzelishvili VZ, Smallwood S, Tower D, Hall RL, Hunt DM, Moyer SA: A single amino acid change in the L-polymerase protein of vesicular stomatitis virus completely abolishes viral mRNA cap methylation. J Virol. 2005 Jun;79(12):7327-37. [PubMed:15919887 ]
- Lockridge O: Genetic variants of human serum cholinesterase influence metabolism of the muscle relaxant succinylcholine. Pharmacol Ther. 1990;47(1):35-60. [PubMed:2195556 ]
- Franklin RB, Zou J, Yu Z, Costello LC: EAAC1 is expressed in rat and human prostate epithelial cells; functions as a high-affinity L-aspartate transporter; and is regulated by prolactin and testosterone. BMC Biochem. 2006 Mar 27;7:10. [PubMed:16566829 ]
- Advani SJ, Hagglund R, Weichselbaum RR, Roizman B: Posttranslational processing of infected cell proteins 0 and 4 of herpes simplex virus 1 is sequential and reflects the subcellular compartment in which the proteins localize. J Virol. 2001 Sep;75(17):7904-12. [PubMed:11483735 ]
- Wang M, Meng Z, Fu J: Synthesis and biodistribution of six novel 99mTc complexes of 2-hydroxybenzaldehyde-amino acid Schiff bases. Appl Radiat Isot. 2006 Feb;64(2):235-40. [PubMed:16309915 ]
- Fisher G, Lopez S, Peterson K, Goff T, Philip I, Gaviria R, Lorenzo N, Tsesarskaia M: Is there a correlation between age and D: -aspartic acid in human knee cartilage? Amino Acids. 2006 Jun 1;. [PubMed:16738792 ]
- Baslow MH: Brain N-acetylaspartate as a molecular water pump and its role in the etiology of Canavan disease: a mechanistic explanation. J Mol Neurosci. 2003;21(3):185-90. [PubMed:14645985 ]
- Shao B, Belaaouaj A, Verlinde CL, Fu X, Heinecke JW: Methionine sulfoxide and proteolytic cleavage contribute to the inactivation of cathepsin G by hypochlorous acid: an oxidative mechanism for regulation of serine proteinases by myeloperoxidase. J Biol Chem. 2005 Aug 12;280(32):29311-21. Epub 2005 Jun 20. [PubMed:15967795 ]
- Rose CH, Thigpen BD, Bofill JA, Cushman J, May WL, Martin JN Jr: Obstetric implications of antepartum corticosteroid therapy for HELLP syndrome. Obstet Gynecol. 2004 Nov;104(5 Pt 1):1011-4. [PubMed:15516393 ]
- Bhende PM, Seaman WT, Delecluse HJ, Kenney SC: BZLF1 activation of the methylated form of the BRLF1 immediate-early promoter is regulated by BZLF1 residue 186. J Virol. 2005 Jun;79(12):7338-48. [PubMed:15919888 ]
- Butterworth RF: Pathophysiology of hepatic encephalopathy: a new look at ammonia. Metab Brain Dis. 2002 Dec;17(4):221-7. [PubMed:12602499 ]
- Elshenawy S, Pinney SE, Stuart T, Doulias PT, Zura G, Parry S, Elovitz MA, Bennett MJ, Bansal A, Strauss JF 3rd, Ischiropoulos H, Simmons RA: The Metabolomic Signature of the Placenta in Spontaneous Preterm Birth. Int J Mol Sci. 2020 Feb 4;21(3). pii: ijms21031043. doi: 10.3390/ijms21031043. [PubMed:32033212 ]
| Show more...
|---|