| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected and Quantified |
|---|
| Creation Date | 2005-11-16 15:48:42 UTC |
|---|
| Update Date | 2023-05-30 20:55:56 UTC |
|---|
| HMDB ID | HMDB0000208 |
|---|
| Secondary Accession Numbers | - HMDB0002812
- HMDB00208
- HMDB0062781
- HMDB02812
- HMDB62781
| Show more accession numbers
|---|
| Metabolite Identification |
|---|
| Common Name | Oxoglutaric acid |
|---|
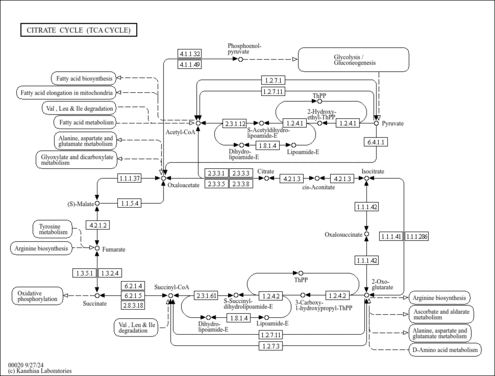
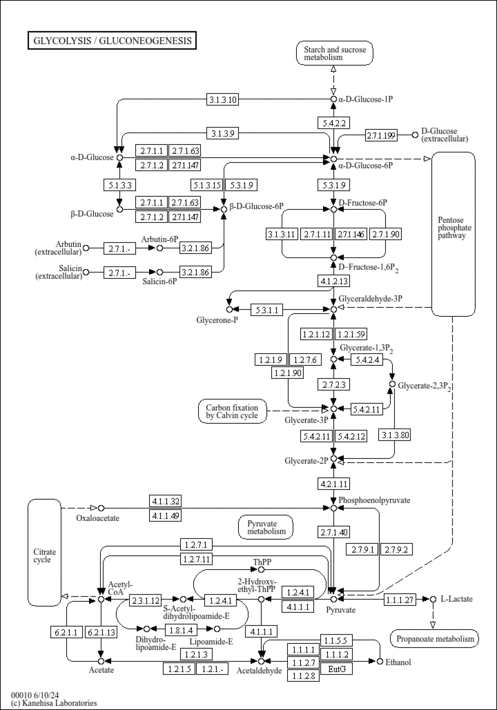
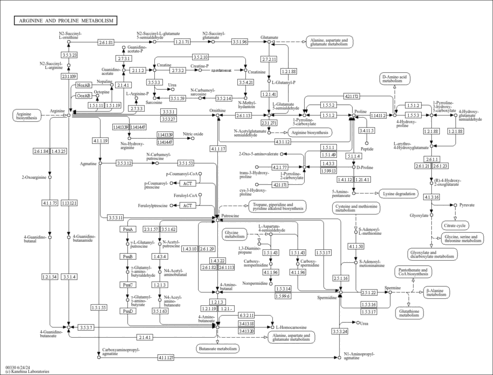
| Description | Oxoglutaric acid, also known as alpha-ketoglutarate, alpha-ketoglutaric acid, AKG, or 2-oxoglutaric acid, is classified as a gamma-keto acid or a gamma-keto acid derivative. gamma-Keto acids are organic compounds containing an aldehyde substituted with a keto group on the C4 carbon atom. alpha-Ketoglutarate is considered to be soluble (in water) and acidic. alpha-Ketoglutarate is a key molecule in the TCA cycle, playing a fundamental role in determining the overall rate of this important metabolic process (PMID: 26759695 ). In the TCA cycle, AKG is decarboxylated to succinyl-CoA and carbon dioxide by AKG dehydrogenase, which functions as a key control point of the TCA cycle. Additionally, AKG can be generated from isocitrate by oxidative decarboxylation catalyzed by the enzyme known as isocitrate dehydrogenase (IDH). In addition to these routes of production, AKG can be produced from glutamate by oxidative deamination via glutamate dehydrogenase, and as a product of pyridoxal phosphate-dependent transamination reactions (mediated by branched-chain amino acid transaminases) in which glutamate is a common amino donor. AKG is a nitrogen scavenger and a source of glutamate and glutamine that stimulates protein synthesis and inhibits protein degradation in muscles. In particular, AKG can decrease protein catabolism and increase protein synthesis to enhance bone tissue formation in skeletal muscles (PMID: 26759695 ). Interestingly, enteric feeding of AKG supplements can significantly increase circulating plasma levels of hormones such as insulin, growth hormone, and insulin-like growth factor-1 (PMID: 26759695 ). It has recently been shown that AKG can extend the lifespan of adult C. elegans by inhibiting ATP synthase and TOR (PMID: 24828042 ). In combination with molecular oxygen, alpha-ketoglutarate is required for the hydroxylation of proline to hydroxyproline in the production of type I collagen. A recent study has shown that alpha-ketoglutarate promotes TH1 differentiation along with the depletion of glutamine thereby favouring Treg (regulatory T-cell) differentiation (PMID: 26420908 ). alpha-Ketoglutarate has been found to be associated with fumarase deficiency, 2-ketoglutarate dehydrogenase complex deficiency, and D-2-hydroxyglutaric aciduria, which are all inborn errors of metabolism (PMID: 8338207 ). Oxoglutaric acid has been found to be a metabolite produced by Corynebacterium and yeast (PMID: 27872963 ) (YMDB). | Read more...
|---|
| Structure | InChI=1S/C5H6O5/c6-3(5(9)10)1-2-4(7)8/h1-2H2,(H,7,8)(H,9,10) |
|---|
| Synonyms | | Value | Source |
|---|
| 2-Ketoglutaric acid | ChEBI | | alpha-Ketoglutaric acid | ChEBI | | 2-Ketoglutarate | Generator | | a-Ketoglutarate | Generator | | a-Ketoglutaric acid | Generator | | alpha-Ketoglutarate | Generator | | Α-ketoglutarate | Generator | | Α-ketoglutaric acid | Generator | | Oxoglutarate | Generator | | 2-oxo-1,5-Pentanedioate | HMDB | | 2-oxo-1,5-Pentanedioic acid | HMDB | | 2-Oxoglutarate | HMDB | | 2-Oxoglutaric acid | HMDB | | 2-Oxopentanedioate | HMDB | | 2-Oxopentanedioic acid | HMDB | | alpha-Oxoglutarate | HMDB | | Oxogluric acid | HMDB | | 2 Oxoglutaric acid | HMDB | | Ketoglutaric acid | HMDB | | alpha Ketoglutaric acid | HMDB | | 2 Ketoglutaric acid | HMDB | | alpha Ketoglutarate | HMDB | | alpha Oxoglutarate | HMDB | | 2 Ketoglutarate | HMDB | | 2 Oxoglutarate | HMDB | | alpha-Oxoglutaric acid | HMDB | | alpha-Oxopentanedioic acid | HMDB | | alpha-Keto-glutaric acid | HMDB | | Α-oxoglutaric acid | HMDB | | Α-oxopentanedioic acid | HMDB | | Α-keto-glutaric acid | HMDB |
| Show more...
|---|
| Chemical Formula | C5H6O5 |
|---|
| Average Molecular Weight | 146.0981 |
|---|
| Monoisotopic Molecular Weight | 146.021523302 |
|---|
| IUPAC Name | 2-oxopentanedioic acid |
|---|
| Traditional Name | oxoglutarate |
|---|
| CAS Registry Number | 328-50-7 |
|---|
| SMILES | OC(=O)CCC(=O)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C5H6O5/c6-3(5(9)10)1-2-4(7)8/h1-2H2,(H,7,8)(H,9,10) |
|---|
| InChI Key | KPGXRSRHYNQIFN-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as gamma-keto acids and derivatives. These are organic compounds containing an aldehyde substituted with a keto group on the C4 carbon atom. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Keto acids and derivatives |
|---|
| Sub Class | Gamma-keto acids and derivatives |
|---|
| Direct Parent | Gamma-keto acids and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Gamma-keto acid
- Short-chain keto acid
- Dicarboxylic acid or derivatives
- Alpha-keto acid
- Alpha-hydroxy ketone
- Ketone
- Carboxylic acid
- Carboxylic acid derivative
- Organic oxygen compound
- Organic oxide
- Hydrocarbon derivative
- Organooxygen compound
- Carbonyl group
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Physiological effect | |
|---|
| Disposition | Biological locationRoute of exposureSourceExogenous- Exogenous (HMDB: HMDB0000208)
Food- Food (HMDB: HMDB0000208)
Animal originHerb and spiceVegetableFruitNutCereal and cereal productPulseGourdCoffee and coffee productSoyDishTeaBaking goodBeverageAquatic originMilk and milk productUnfermented milk- Milk (Cow) (FooDB: FOOD00618)
- Cow milk, pasteurized, vitamin A + D added, 0% fat (FooDB: FOOD00889)
- Cow milk, pasteurized, vitamin A + D added, 1% fat (FooDB: FOOD00890)
- Cow milk, pasteurized, vitamin A + D added, 2% fat (FooDB: FOOD00891)
- Cow milk, pasteurized, vitamin D added, 3.25% fat (FooDB: FOOD00892)
Cocoa and cocoa product
Endogenous |
|---|
| Process | Naturally occurring processBiological processBiochemical pathwayMetabolic pathwayDisease pathway- 2-Hydroxyglutric Aciduria (D and L Form) (HMDB: HMDB0000208)
- 2-Ketoglutarate Dehydrogenase Complex Deficiency (HMDB: HMDB0000208)
- Pyruvate Carboxylase Deficiency (HMDB: HMDB0000208)
- Fumarase Deficiency (HMDB: HMDB0000208)
- Pyruvate Dehydrogenase Deficiency (E3) (HMDB: HMDB0000208)
- Cystinosis, Ocular Nonnephropathic (HMDB: HMDB0000208)
- The Oncogenic Action of Succinate (PathBank: SMP0002292)
- Glutaminolysis and Cancer (PathBank: SMP0108773)
- 2-Methyl-3-hydroxybutryl-CoA Dehydrogenase Deficiency (PathBank: SMP0120439)
- 4-Hydroxybutyric Aciduria/Succinic Semialdehyde Dehydrogenase Deficiency (PathBank: SMP0120445)
- Arginine: Glycine Amidinotransferase Deficiency (AGAT Deficiency) (PathBank: SMP0120454)
- GABA-Transaminase Deficiency (PathBank: SMP0120478)
- Sarcosinemia (PathBank: SMP0120521)
- Methylmalonate Semialdehyde Dehydrogenase Deficiency (PathBank: SMP0120523)
- Refsum Disease (PathBank: SMP0120529)
- Creatine Deficiency, Guanidinoacetate Methyltransferase Deficiency (PathBank: SMP0120569)
- Isovaleric Acidemia (PathBank: SMP0120589)
- Fructose-1,6-diphosphatase Deficiency (PathBank: SMP0120626)
- Succinic Semialdehyde Dehydrogenase Deficiency (PathBank: SMP0120631)
- Pyridoxine Dependency with Seizures (PathBank: SMP0120635)
- 3-Methylglutaconic Aciduria Type I (PathBank: SMP0120663)
- Argininemia (PathBank: SMP0120676)
- Argininosuccinic Aciduria (PathBank: SMP0120677)
- Beta-Ketothiolase Deficiency (PathBank: SMP0120679)
- Canavan Disease (PathBank: SMP0120681)
- Carbamoyl Phosphate Synthetase Deficiency (PathBank: SMP0120682)
- Dihydropyrimidine Dehydrogenase Deficiency (DHPD) (PathBank: SMP0120694)
- Glutaric Aciduria Type I (PathBank: SMP0120704)
- Hyperprolinemia Type I (PathBank: SMP0120719)
- Hypoacetylaspartia (PathBank: SMP0120722)
- Tyrosinemia Type 3 (TYRO3) (PathBank: SMP0120731)
- Ornithine Transcarbamylase Deficiency (OTC Deficiency) (PathBank: SMP0120762)
- Tyrosinemia, Transient, of the Newborn (PathBank: SMP0120779)
- Beta-Mercaptolactate-Cysteine Disulfiduria (PathBank: SMP0120783)
- Hyperornithinemia with Gyrate Atrophy (HOGA) (PathBank: SMP0120789)
- 3-Hydroxyisobutyric Aciduria (PathBank: SMP0120806)
- Pyruvate Dehydrogenase Deficiency (E2) (PathBank: SMP0120834)
- Glycogenosis, Type IC (PathBank: SMP0120857)
- Glycogenosis, Type IA. Von Gierke Disease (PathBank: SMP0120864)
- Hyperprolinemia Type II (PathBank: SMP0120501)
- The Oncogenic Action of 2-Hydroxyglutarate (PathBank: SMP0002291)
- 3-Methylcrotonyl-CoA Carboxylase Deficiency Type I (PathBank: SMP0120441)
- 3-Methylglutaconic Aciduria Type IV (PathBank: SMP0120444)
- Tyrosinemia Type 2 (or Richner-Hanhart Syndrome) (PathBank: SMP0120513)
- Methylmalonic Aciduria (PathBank: SMP0120525)
- Non-Ketotic Hyperglycinemia (PathBank: SMP0120534)
- Prolinemia Type II (PathBank: SMP0120539)
- Phenylketonuria (PathBank: SMP0120540)
- Prolidase Deficiency (PD) (PathBank: SMP0120541)
- Primary Hyperoxaluria Type I (PathBank: SMP0120543)
- Carnosinuria, Carnosinemia (PathBank: SMP0120558)
- L-Arginine:Glycine Amidinotransferase Deficiency (PathBank: SMP0120572)
- Congenital Lactic Acidosis (PathBank: SMP0120610)
- 3-Hydroxy-3-methylglutaryl-CoA Lyase Deficiency (PathBank: SMP0120661)
- 3-Methylglutaconic Aciduria Type III (PathBank: SMP0120664)
- Hawkinsinuria (PathBank: SMP0120711)
- Lactic Acidemia (PathBank: SMP0120724)
- Tyrosinemia Type I (PathBank: SMP0120729)
- Methylmalonic Aciduria Due to Cobalamin-Related Disorders (PathBank: SMP0120748)
- Hyperornithinemia-Hyperammonemia-Homocitrullinuria [HHH-syndrome] (PathBank: SMP0120790)
- Hyperlysinemia I, Familial (PathBank: SMP0120811)
- Hyperlysinemia II or Saccharopinuria (PathBank: SMP0120812)
- Phosphoenolpyruvate Carboxykinase Deficiency 1 (PEPCK1) (PathBank: SMP0120843)
- Triosephosphate Isomerase Deficiency (PathBank: SMP0120846)
- 2-Aminoadipic 2-Oxoadipic Aciduria (PathBank: SMP0120870)
- 3-Phosphoglycerate Dehydrogenase Deficiency (PathBank: SMP0120872)
- Alkaptonuria (PathBank: SMP0120453)
- Citrullinemia Type I (PathBank: SMP0120466)
- Glycogen Storage Disease Type 1A (GSD1A) or Von Gierke Disease (PathBank: SMP0120488)
- Homocarnosinosis (PathBank: SMP0120494)
- Maple Syrup Urine Disease (PathBank: SMP0120516)
- Propionic Acidemia (PathBank: SMP0120537)
- Ornithine Aminotransferase Deficiency (OAT Deficiency) (PathBank: SMP0120538)
- Dopamine beta-Hydroxylase Deficiency (PathBank: SMP0120563)
- Isobutyryl-CoA Dehydrogenase Deficiency (PathBank: SMP0120588)
- Glycogenosis, Type IB (PathBank: SMP0120637)
- Dimethylglycine Dehydrogenase Deficiency (PathBank: SMP0120695)
- Guanidinoacetate Methyltransferase Deficiency (GAMT Deficiency) (PathBank: SMP0120710)
- Isovaleric Aciduria (PathBank: SMP0120723)
- Malonic Aciduria (PathBank: SMP0120734)
- Saccharopinuria/Hyperlysinemia II (PathBank: SMP0120746)
- Hyperglycinemia, Non-Ketotic (PathBank: SMP0120770)
- Malonyl-CoA Decarboxylase Deficiency (PathBank: SMP0120786)
- Monoamine Oxidase-A Deficiency (MAO-A) (PathBank: SMP0120817)
- Mitochondrial Complex II Deficiency (PathBank: SMP0120831)
- The Oncogenic Action of Fumarate (PathBank: SMP0002295)
- Hyperinsulinism-Hyperammonemia Syndrome (PathBank: SMP0120496)
- 3-Hydroxyisobutyric Acid Dehydrogenase Deficiency (PathBank: SMP0120586)
- Cystinosis, Ocular Nonnephropathic (PathBank: SMP0120654)
- 2-Methyl-3-hydroxybutryl-CoA Dehydrogenase Deficiency (PathBank: SMP0000137)
- Hyperornithinemia-Hyperammonemia-Homocitrullinuria [HHH-syndrome] (PathBank: SMP0000506)
- The Oncogenic Action of L-2-Hydroxyglutarate in Hydroxygluaricaciduria (PathBank: SMP0002358)
- The Oncogenic Action of D-2-Hydroxyglutarate in Hydroxygluaricaciduria (PathBank: SMP0002359)
Protein pathwayDrug action pathway |
|---|
| Role | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | 115.5 °C | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | 541.5 mg/mL | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Experimental Chromatographic Properties | Experimental Collision Cross Sections |
|---|
| Predicted Molecular Properties | | Show more...
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Kovats Retention IndicesUnderivatizedDerivatized| Derivative Name / Structure | SMILES | Kovats RI Value | Column Type | Reference |
|---|
| Oxoglutaric acid,1TMS,isomer #1 | C[Si](C)(C)OC(=O)CCC(=O)C(=O)O | 1427.4 | Semi standard non polar | 33892256 | | Oxoglutaric acid,1TMS,isomer #2 | C[Si](C)(C)OC(=O)C(=O)CCC(=O)O | 1400.3 | Semi standard non polar | 33892256 | | Oxoglutaric acid,1TMS,isomer #3 | C[Si](C)(C)OC(=CCC(=O)O)C(=O)O | 1518.3 | Semi standard non polar | 33892256 | | Oxoglutaric acid,2TMS,isomer #1 | C[Si](C)(C)OC(=O)CCC(=O)C(=O)O[Si](C)(C)C | 1545.0 | Semi standard non polar | 33892256 | | Oxoglutaric acid,2TMS,isomer #2 | C[Si](C)(C)OC(=O)CC=C(O[Si](C)(C)C)C(=O)O | 1594.5 | Semi standard non polar | 33892256 | | Oxoglutaric acid,2TMS,isomer #3 | C[Si](C)(C)OC(=O)C(=CCC(=O)O)O[Si](C)(C)C | 1557.7 | Semi standard non polar | 33892256 | | Oxoglutaric acid,3TMS,isomer #1 | C[Si](C)(C)OC(=O)CC=C(O[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1617.1 | Semi standard non polar | 33892256 | | Oxoglutaric acid,3TMS,isomer #1 | C[Si](C)(C)OC(=O)CC=C(O[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1573.4 | Standard non polar | 33892256 | | Oxoglutaric acid,3TMS,isomer #1 | C[Si](C)(C)OC(=O)CC=C(O[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1600.2 | Standard polar | 33892256 | | Oxoglutaric acid,1TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)CCC(=O)C(=O)O | 1706.3 | Semi standard non polar | 33892256 | | Oxoglutaric acid,1TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC(=O)C(=O)CCC(=O)O | 1675.5 | Semi standard non polar | 33892256 | | Oxoglutaric acid,1TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)OC(=CCC(=O)O)C(=O)O | 1770.5 | Semi standard non polar | 33892256 | | Oxoglutaric acid,2TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)CCC(=O)C(=O)O[Si](C)(C)C(C)(C)C | 1988.0 | Semi standard non polar | 33892256 | | Oxoglutaric acid,2TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC(=O)CC=C(O[Si](C)(C)C(C)(C)C)C(=O)O | 2069.5 | Semi standard non polar | 33892256 | | Oxoglutaric acid,2TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)OC(=O)C(=CCC(=O)O)O[Si](C)(C)C(C)(C)C | 2025.1 | Semi standard non polar | 33892256 | | Oxoglutaric acid,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)CC=C(O[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2255.6 | Semi standard non polar | 33892256 | | Oxoglutaric acid,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)CC=C(O[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2128.5 | Standard non polar | 33892256 | | Oxoglutaric acid,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)CC=C(O[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2045.3 | Standard polar | 33892256 |
| Show more...
|---|
| Spectra |
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental GC-MS | GC-MS Spectrum - Oxoglutaric acid GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (2 TMS) | splash10-000b-3910000000-4097b7eb52c2910a4277 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoglutaric acid GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (Non-derivatized) | splash10-0002-1910000000-3277227b9c4da4baa1f7 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoglutaric acid GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (2 TMS; 1 MEOX) | splash10-00di-9300000000-974bce546caa4fbe6b0f | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoglutaric acid GC-MS (1 MEOX; 2 TMS) | splash10-000i-9710000000-d0aea11e947b042571f9 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoglutaric acid GC-MS (1 MEOX; 2 TMS) | splash10-052s-4920000000-bf7ed92c68fd4c8dbb6b | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoglutaric acid GC-EI-TOF (Non-derivatized) | splash10-000b-3910000000-4097b7eb52c2910a4277 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoglutaric acid GC-EI-TOF (Non-derivatized) | splash10-0002-1910000000-3277227b9c4da4baa1f7 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoglutaric acid GC-EI-TOF (Non-derivatized) | splash10-00di-9300000000-974bce546caa4fbe6b0f | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoglutaric acid GC-MS (Non-derivatized) | splash10-000i-9710000000-d0aea11e947b042571f9 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoglutaric acid GC-MS (Non-derivatized) | splash10-052s-4920000000-bf7ed92c68fd4c8dbb6b | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoglutaric acid GC-EI-TOF (Non-derivatized) | splash10-000b-2900000000-e514f7afea2490bbe9ed | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoglutaric acid GC-MS (Non-derivatized) - 70eV, Positive | splash10-0kfx-9200000000-0ee77a39e9862659f8a7 | 2016-09-22 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoglutaric acid GC-MS (2 TMS) - 70eV, Positive | splash10-00di-9730000000-5e17bd8cfd944963c79f | 2017-10-06 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoglutaric acid GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoglutaric acid GC-MS (TMS_1_1) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoglutaric acid GC-MS (TMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoglutaric acid GC-MS (TMS_1_3) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoglutaric acid GC-MS (TMS_2_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoglutaric acid GC-MS (TMS_2_3) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoglutaric acid GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoglutaric acid GC-MS (TBDMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoglutaric acid GC-MS (TBDMS_1_3) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoglutaric acid GC-MS (TBDMS_2_1) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoglutaric acid GC-MS (TBDMS_2_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoglutaric acid GC-MS (TBDMS_2_3) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-0fba-9200000000-4e367c66e94c51eb1a1b | 2014-09-20 | Not Available | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoglutaric acid Quattro_QQQ 10V, Negative-QTOF (Annotated) | splash10-0zfr-6900000000-7fc343a243ce5d2a4f39 | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoglutaric acid Quattro_QQQ 25V, Negative-QTOF (Annotated) | splash10-0a4i-9200000000-1a7af9946604baa8b3d2 | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoglutaric acid Quattro_QQQ 40V, Negative-QTOF (Annotated) | splash10-0a4l-9800000000-30495b2afdd3526854e7 | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoglutaric acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-0002-0925200000-af33449c9638276466c2 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoglutaric acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-0a4i-9000000000-6e43f767847790112b65 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoglutaric acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-03di-0900000000-4cacc41705b7b385e91e | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoglutaric acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-0udi-0900000000-1367eeeb6ad64c67a0aa | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoglutaric acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-000t-0915051100-91cda631ee54800322b8 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoglutaric acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-0a4i-9000000000-4f842c3673925d29c402 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoglutaric acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-0udi-0900000000-5e751351656f6eb62de3 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoglutaric acid LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-001i-0910000000-7729faa24733fbad77b7 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoglutaric acid LC-ESI-QQ (API3000, Applied Biosystems) 10V, Negative-QTOF | splash10-0002-0900000000-c3c62f83bff99f44f7ab | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoglutaric acid LC-ESI-QQ (API3000, Applied Biosystems) 20V, Negative-QTOF | splash10-0zfr-6900000000-b6d8379db3e3f1480b14 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoglutaric acid LC-ESI-QQ (API3000, Applied Biosystems) 30V, Negative-QTOF | splash10-0a4i-9300000000-4fde39bb93cde5a27f23 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoglutaric acid LC-ESI-QQ (API3000, Applied Biosystems) 40V, Negative-QTOF | splash10-00i0-9000000000-4343832f3a55842423a9 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoglutaric acid 10V, Positive-QTOF | splash10-004i-2900000000-5aa3d6d620db1d733a65 | 2015-05-26 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoglutaric acid 20V, Positive-QTOF | splash10-0m30-7900000000-fc77f69ad43f915bf23e | 2015-05-26 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoglutaric acid 40V, Positive-QTOF | splash10-0a6r-9000000000-c597f1f5e416037aa82d | 2015-05-26 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoglutaric acid 10V, Positive-QTOF | splash10-004i-2900000000-5aa3d6d620db1d733a65 | 2015-05-26 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoglutaric acid 20V, Positive-QTOF | splash10-0m30-7900000000-fc77f69ad43f915bf23e | 2015-05-26 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoglutaric acid 40V, Positive-QTOF | splash10-0a6r-9000000000-c597f1f5e416037aa82d | 2015-05-26 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoglutaric acid 10V, Negative-QTOF | splash10-0002-1900000000-f2b463076a150a514f0c | 2015-05-27 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoglutaric acid 20V, Negative-QTOF | splash10-0f9t-8900000000-b2d3fa8d7ab01a220d9a | 2015-05-27 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoglutaric acid 40V, Negative-QTOF | splash10-053r-9100000000-554bfbd6839bc376cba0 | 2015-05-27 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoglutaric acid 10V, Negative-QTOF | splash10-0002-1900000000-f2b463076a150a514f0c | 2015-05-27 | Wishart Lab | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Experimental 1D NMR | 1H NMR Spectrum (1D, 600 MHz, H2O, experimental) | 2012-12-04 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Experimental 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, H2O, experimental) | 2012-12-05 | Wishart Lab | View Spectrum |
IR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M-H]-) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+H]+) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+Na]+) | 2023-02-03 | FELIX lab | View Spectrum |
| Show more...
|---|
| Biological Properties |
|---|
| Cellular Locations | - Extracellular
- Mitochondria
- Endoplasmic reticulum
- Peroxisome
|
|---|
| Biospecimen Locations | - Blood
- Breast Milk
- Cellular Cytoplasm
- Cerebrospinal Fluid (CSF)
- Feces
- Saliva
- Urine
|
|---|
| Tissue Locations | |
|---|
| Pathways | |
|---|
| Normal Concentrations |
|---|
| |
| Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 7.0 (0.0-23.0) uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 8.6 +/- 2.6 uM | Children (1-13 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 9.3 +/- 2.3 uM | Children (1-13 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 8.9 +/- 2.7 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 4-13 uM | Not Specified | Not Specified | Normal | | details | | Blood | Detected and Quantified | 20.7 +/- 2.5 uM | Newborn (0-30 days old) | Both | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Breast Milk | Detected and Quantified | 41.0 +/- 16.1 uM | Adult (>18 years old) | Female | Normal | | details | | Cellular Cytoplasm | Detected and Quantified | 320 (290-350) uM | Adult (>18 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 5 +/- 4 uM | Adult (>18 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 2.1 +/- 0.7 uM | Children (1-13 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 8.9 +/- 3.4 uM | Adult (>18 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 16-Feb uM | Not Specified | Not Specified | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 2.0 +/- 2.0 uM | Adult (>18 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 4.80 (3.40-6.20) uM | Children (1-13 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Infant (0-1 year old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 5.27 +/- 3.61 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 1.53 +/- 1.18 uM | Adult (>18 years old) | Female | Normal | | details | | Saliva | Detected and Quantified | 1.64 +/- 3.47 uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 2.65 +/- 1.07 uM | Adult (>18 years old) | Female | Normal | | details | | Saliva | Detected and Quantified | 2.65 +/- 1.07 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Saliva | Detected and Quantified | 3.78 +/- 2.33 uM | Adult (>18 years old) | Female | Normal | | details | | Urine | Detected and Quantified | <110 umol/mmol creatinine | Newborn (0-30 days old) | Female | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 32.164 +/- 9.964 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 30.79-398.2 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 12-20 umol/mmol creatinine | Adult (>18 years old) | Female | Normal | | details | | Urine | Detected and Quantified | 90.83(45.64-331.1) umol/mmol creatinine | Newborn (0-30 days old) | Female | Normal | | details | | Urine | Detected and Quantified | <150 umol/mmol creatinine | Adult (>18 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 104.8(30.21-413.8) umol/mmol creatinine | Newborn (0-30 days old) | Male | Normal | | details | | Urine | Detected and Quantified | 2.87 (0.18-14.3) umol/mmol creatinine | Adult (>18 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 2.98 (0.42-18.3) umol/mmol creatinine | Adult (>18 years old) | Female | Normal | | details | | Urine | Detected and Quantified | 23.6 umol/mmol creatinine | Infant (0-1 year old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 29.893 +/- 15.594 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | <60 umol/mmol creatinine | Not Specified | Not Specified | Normal | | details | | Urine | Detected and Quantified | <75 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | <95 umol/mmol creatinine | Children (1-13 years old) | Female | Normal | | details | | Urine | Detected and Quantified | <115 umol/mmol creatinine | Children (1-13 years old) | Female | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | <350 umol/mmol creatinine | Infant (0-1 year old) | Male | Normal | | details | | Urine | Detected and Quantified | 0-25.3 umol/mmol creatinine | Infant (0-1 year old) | Male | Normal | | details | | Urine | Detected and Quantified | 14.8-64.7 umol/mmol creatinine | Infant (0-1 year old) | Female | Normal | | details | | Urine | Detected and Quantified | 4.3-36.3 umol/mmol creatinine | Infant (0-1 year old) | Male | Normal | | details | | Urine | Detected and Quantified | 7.5-28.9 umol/mmol creatinine | Infant (0-1 year old) | Female | Normal | | details | | Urine | Detected and Quantified | 8.90-17.12 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 4.2 +/- 1.97 umol/mmol creatinine | Children (1-13 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 24 (4-74) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 150.5 (10.0-836.3) umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 236.9 (8.0-631.4) umol/mmol creatinine | Infant (0-1 year old) | Both | Normal | | details | | Urine | Detected and Quantified | 21.4 (4.9-123.8) umol/mmol creatinine | Children (1-13 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 25.6 (10.2-146.0) umol/mmol creatinine | Adolescent (13-18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 15.477-154.773 umol/mmol creatinine | Infant (0-1 year old) | Both | Normal | | details | | Urine | Detected and Quantified | 7-11 umol/mmol creatinine | Adult (>18 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 4.8 (2.0-17.0) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 2.2-8.2 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 162.32 (71.28-284.43) umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 0.083 +/- 0.002 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 1.548-39.467 umol/mmol creatinine | Children (1 - 13 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 43.46 +/- 53.46 umol/mmol creatinine | Infant (0-1 year old) | Both | Normal | | details | | Urine | Detected and Quantified | <79 umol/mmol creatinine | Children (1-13 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | > 0.658 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 160-590 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 3.4 (1.4-12.5) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details |
| Show more...
|---|
| Abnormal Concentrations |
|---|
| |
| Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Schizophrenia | | details | | Blood | Detected and Quantified | 17-37 uM | Children (1-13 years old) | Both | 2-Ketoglutarate dehydrogenase complex deficiency | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Schizophrenia | | details | | Cellular Cytoplasm | Detected and Quantified | 150 (144-156) uM | Adult (>18 years old) | Both | Anoxia | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 3.600-12.800 uM | Children (1-13 years old) | Both | 2-Ketoglutarate dehydrogenase complex deficiency | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal Cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Immunoglobulin A nephropathy (IgAN) progressor | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Urine | Detected and Quantified | 1374 umol/mmol creatinine | Newborn (0-30 days old) | Female | Deafness, Onychodystrophy, Osteodystrophy, Mental Retardation, and Seizures Syndrome | | details | | Urine | Detected and Quantified | 28.849 +/- 8.834 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Eosinophilic esophagitis | | details | | Urine | Detected and Quantified | 39.88 +/- 30.024 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Gastroesophageal reflux disease | | details | | Urine | Detected and Quantified | 30-1600 umol/mmol creatinine | Infant (0-1 year old) | Male | 2-Ketoglutarate dehydrogenase complex deficiency | | details | | Urine | Detected and Quantified | 24.882 +/- 26.016 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Eosinophilic esophagitis | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Schizophrenia | | details | | Urine | Detected and Quantified | 100-166 umol/mmol creatinine | Adult (>18 years old) | Both | Hyperinsulinism-hyperammonemia syndrome | | details | | Urine | Detected and Quantified | 149-247 umol/mmol creatinine | Children (1-13 years old) | Female | Hyperinsulinism-hyperammonemia syndrome | | details | | Urine | Detected and Quantified | 15-107 umol/mmol creatinine | Children (1-13 years old) | Female | Hyperinsulinism-hyperammonemia syndrome | | details | | Urine | Detected and Quantified | 1172.1 umol/mmol creatinine | Infant (0-1 year old) | Male | Fumarase deficiency | | details | | Urine | Detected and Quantified | 385.200-1367.300 umol/mmol creatinine | Infant (0-1 year old) | Male | Fumarase deficiency | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Bladder cancer | | details | | Urine | Detected and Quantified | 673.261-2970.0885 umol/mmol creatinine | Infant (0-1 year old) | Both | Amish lethal microcephaly | | details | | Urine | Detected and Quantified | 1119.294 umol/mmol creatinine | Infant (0-1 year old) | Male | Dihydrolipoamide Dehydrogenase Deficiency | | details | | Urine | Detected and Quantified | 26.667 umol/mmol creatinine | Children (1 - 13 years old) | Male | D-2-Hydroxyglutaric aciduria | | details | | Urine | Detected and Quantified | 46.432 umol/mmol creatinine | Children (1 - 13 years old) | Male | D-2-Hydroxyglutaric aciduria | | details | | Urine | Detected and Quantified | 193.466 umol/mmol creatinine | Children (1 - 13 years old) | Male | D-2-Hydroxyglutaric aciduria | | details | | Urine | Detected and Quantified | 6224 umol/mmol creatinine | Children (1-13 years old) | Male | Lipoyltransferase 1 Deficiency | | details | | Urine | Detected and Quantified | 101.0 (50.0-152.0) umol/mmol creatinine | Adult (>18 years old) | Both | 2-Ketoglutarate dehydrogenase complex deficiency | | details | | Urine | Detected and Quantified | 875.0 (150.0-1600.0) umol/mmol creatinine | Children (1-13 years old) | Both | 2-Ketoglutarate Dehydrogenase Complex Deficiency | | details |
| Show more...
|---|
| Associated Disorders and Diseases |
|---|
| Disease References | | Schizophrenia |
|---|
- Yang J, Chen T, Sun L, Zhao Z, Qi X, Zhou K, Cao Y, Wang X, Qiu Y, Su M, Zhao A, Wang P, Yang P, Wu J, Feng G, He L, Jia W, Wan C: Potential metabolite markers of schizophrenia. Mol Psychiatry. 2013 Jan;18(1):67-78. doi: 10.1038/mp.2011.131. Epub 2011 Oct 25. [PubMed:22024767 ]
- Xuan J, Pan G, Qiu Y, Yang L, Su M, Liu Y, Chen J, Feng G, Fang Y, Jia W, Xing Q, He L: Metabolomic profiling to identify potential serum biomarkers for schizophrenia and risperidone action. J Proteome Res. 2011 Dec 2;10(12):5433-43. doi: 10.1021/pr2006796. Epub 2011 Nov 8. [PubMed:22007635 ]
- Cai HL, Li HD, Yan XZ, Sun B, Zhang Q, Yan M, Zhang WY, Jiang P, Zhu RH, Liu YP, Fang PF, Xu P, Yuan HY, Zhang XH, Hu L, Yang W, Ye HS: Metabolomic analysis of biochemical changes in the plasma and urine of first-episode neuroleptic-naive schizophrenia patients after treatment with risperidone. J Proteome Res. 2012 Aug 3;11(8):4338-50. doi: 10.1021/pr300459d. Epub 2012 Jul 26. [PubMed:22800120 ]
| | Anoxia |
|---|
- Zupke C, Sinskey AJ, Stephanopoulos G: Intracellular flux analysis applied to the effect of dissolved oxygen on hybridomas. Appl Microbiol Biotechnol. 1995 Dec;44(1-2):27-36. [PubMed:8579834 ]
| | Colorectal cancer |
|---|
- Brown DG, Rao S, Weir TL, O'Malia J, Bazan M, Brown RJ, Ryan EP: Metabolomics and metabolic pathway networks from human colorectal cancers, adjacent mucosa, and stool. Cancer Metab. 2016 Jun 6;4:11. doi: 10.1186/s40170-016-0151-y. eCollection 2016. [PubMed:27275383 ]
| | 2-Ketoglutarate dehydrogenase complex deficiency |
|---|
- Bonnefont JP, Chretien D, Rustin P, Robinson B, Vassault A, Aupetit J, Charpentier C, Rabier D, Saudubray JM, Munnich A: Alpha-ketoglutarate dehydrogenase deficiency presenting as congenital lactic acidosis. J Pediatr. 1992 Aug;121(2):255-8. [PubMed:1640293 ]
- Kohlschutter A, Behbehani A, Langenbeck U, Albani M, Heidemann P, Hoffmann G, Kleineke J, Lehnert W, Wendel U: A familial progressive neurodegenerative disease with 2-oxoglutaric aciduria. Eur J Pediatr. 1982 Feb;138(1):32-7. [PubMed:7075624 ]
- G.Frauendienst-Egger, Friedrich K. Trefz (2017). MetaGene: Metabolic & Genetic Information Center (MIC: http://www.metagene.de). METAGENE consortium.
| | Eosinophilic esophagitis |
|---|
- Slae, M., Huynh, H., Wishart, D.S. (2014). Analysis of 30 normal pediatric urine samples via NMR spectroscopy (unpublished work). NA.
| | D-2-hydroxyglutaric aciduria |
|---|
- Chalmers RA, Lawson AM, Watts RW, Tavill AS, Kamerling JP, Hey E, Ogilvie D: D-2-hydroxyglutaric aciduria: case report and biochemical studies. J Inherit Metab Dis. 1980;3(1):11-5. [PubMed:6774165 ]
| | Hyperinsulinism-hyperammonemia syndrome |
|---|
- Meissner T, Mayatepek E, Kinner M, Santer R: Urinary alpha-ketoglutarate is elevated in patients with hyperinsulinism-hyperammonemia syndrome. Clin Chim Acta. 2004 Mar;341(1-2):23-6. doi: 10.1016/j.cccn.2003.10.023. [PubMed:14967154 ]
| | Fumarase deficiency |
|---|
- Tregoning S, Salter W, Thorburn DR, Durkie M, Panayi M, Wu JY, Easterbrook A, Coman DJ: Fumarase deficiency in dichorionic diamniotic twins. Twin Res Hum Genet. 2013 Dec;16(6):1117-20. doi: 10.1017/thg.2013.72. Epub 2013 Nov 4. [PubMed:24182348 ]
| | Amish lethal microcephaly |
|---|
- Kelley RI, Robinson D, Puffenberger EG, Strauss KA, Morton DH: Amish lethal microcephaly: a new metabolic disorder with severe congenital microcephaly and 2-ketoglutaric aciduria. Am J Med Genet. 2002 Nov 1;112(4):318-26. doi: 10.1002/ajmg.10529. [PubMed:12376931 ]
| | Deafness, Onychodystrophy, Osteodystrophy, Mental Retardation, and Seizures Syndrome |
|---|
- James AW, Miranda SG, Culver K, Hall BD, Golabi M: DOOR syndrome: clinical report, literature review and discussion of natural history. Am J Med Genet A. 2007 Dec 1;143A(23):2821-31. doi: 10.1002/ajmg.a.32054. [PubMed:17994565 ]
| | Dihydrolipoamide Dehydrogenase Deficiency |
|---|
- Kuhara T, Shinka T, Inoue Y, Matsumoto M, Yoshino M, Sakaguchi Y, Matsumoto I: Studies of urinary organic acid profiles of a patient with dihydrolipoyl dehydrogenase deficiency. Clin Chim Acta. 1983 Sep 30;133(2):133-40. [PubMed:6688766 ]
| | Lipoyltransferase 1 Deficiency |
|---|
- Soreze Y, Boutron A, Habarou F, Barnerias C, Nonnenmacher L, Delpech H, Mamoune A, Chretien D, Hubert L, Bole-Feysot C, Nitschke P, Correia I, Sardet C, Boddaert N, Hamel Y, Delahodde A, Ottolenghi C, de Lonlay P: Mutations in human lipoyltransferase gene LIPT1 cause a Leigh disease with secondary deficiency for pyruvate and alpha-ketoglutarate dehydrogenase. Orphanet J Rare Dis. 2013 Dec 17;8:192. doi: 10.1186/1750-1172-8-192. [PubMed:24341803 ]
|
|
|---|
| Associated OMIM IDs | - 181500 (Schizophrenia)
- 114500 (Colorectal cancer)
- 203740 (2-Ketoglutarate dehydrogenase complex deficiency)
- 610247 (Eosinophilic esophagitis)
- 600721 (D-2-hydroxyglutaric aciduria)
- 606762 (Hyperinsulinism-hyperammonemia syndrome)
- 606812 (Fumarase deficiency)
- 607196 (Amish lethal microcephaly)
- 220500 (Deafness, Onychodystrophy, Osteodystrophy, Mental Retardation, and Seizures Syndrome)
- 246900 (Dihydrolipoamide Dehydrogenase Deficiency)
- 616299 (Lipoyltransferase 1 Deficiency)
|
|---|
| External Links |
|---|
| DrugBank ID | DB08845 |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FooDB ID | FDB003361 |
|---|
| KNApSAcK ID | C00000769 |
|---|
| Chemspider ID | 50 |
|---|
| KEGG Compound ID | C00026 |
|---|
| BioCyc ID | 2-KETOGLUTARATE |
|---|
| BiGG ID | 33565 |
|---|
| Wikipedia Link | Alpha-Ketoglutaric_acid |
|---|
| METLIN ID | 5218 |
|---|
| PubChem Compound | 51 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 30915 |
|---|
| Food Biomarker Ontology | Not Available |
|---|
| VMH ID | AKG |
|---|
| MarkerDB ID | MDB00000099 |
|---|
| Good Scents ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Tanaka, Katsunobu; kimura, Kazu; Yamaguchi, Ken. Fermentative production of a-oxoglutaric acid. U.S. (1973), 4 pp. |
|---|
| Material Safety Data Sheet (MSDS) | Not Available |
|---|
| General References | - Nicholson JK, Foxall PJ, Spraul M, Farrant RD, Lindon JC: 750 MHz 1H and 1H-13C NMR spectroscopy of human blood plasma. Anal Chem. 1995 Mar 1;67(5):793-811. [PubMed:7762816 ]
- Sweatman BC, Farrant RD, Holmes E, Ghauri FY, Nicholson JK, Lindon JC: 600 MHz 1H-NMR spectroscopy of human cerebrospinal fluid: effects of sample manipulation and assignment of resonances. J Pharm Biomed Anal. 1993 Aug;11(8):651-64. [PubMed:8257730 ]
- Lee SH, Kim SO, Chung BC: Gas chromatographic-mass spectrometric determination of urinary oxoacids using O-(2,3,4,5,6-pentafluorobenzyl)oxime-trimethylsilyl ester derivatization and cation-exchange chromatography. J Chromatogr B Biomed Sci Appl. 1998 Nov 20;719(1-2):1-7. [PubMed:9869358 ]
- Guneral F, Bachmann C: Age-related reference values for urinary organic acids in a healthy Turkish pediatric population. Clin Chem. 1994 Jun;40(6):862-6. [PubMed:8087979 ]
- Ohdoi C, Nyhan WL, Kuhara T: Chemical diagnosis of Lesch-Nyhan syndrome using gas chromatography-mass spectrometry detection. J Chromatogr B Analyt Technol Biomed Life Sci. 2003 Jul 15;792(1):123-30. [PubMed:12829005 ]
- Zupke C, Sinskey AJ, Stephanopoulos G: Intracellular flux analysis applied to the effect of dissolved oxygen on hybridomas. Appl Microbiol Biotechnol. 1995 Dec;44(1-2):27-36. [PubMed:8579834 ]
- Bales JR, Higham DP, Howe I, Nicholson JK, Sadler PJ: Use of high-resolution proton nuclear magnetic resonance spectroscopy for rapid multi-component analysis of urine. Clin Chem. 1984 Mar;30(3):426-32. [PubMed:6321058 ]
- Stromme JH, Borud O, Moe PJ: Fatal lactic acidosis in a newborn attributable to a congenital defect of pyruvate dehydrogenase. Pediatr Res. 1976 Jan;10(1):62-6. [PubMed:813176 ]
- Rocchiccioli F, Leroux JP, Cartier PH: Microdetermination of 2-ketoglutaric acid in plasma and cerebrospinal fluid by capillary gas chromatography mass spectrometry; application to pediatrics. Biomed Mass Spectrom. 1984 Jan;11(1):24-8. [PubMed:6704500 ]
- Barbas C, Garcia A, de Miguel L, Simo C: Evaluation of filter paper collection of urine samples for detection and measurement of organic acidurias by capillary electrophoresis. J Chromatogr B Analyt Technol Biomed Life Sci. 2002 Nov 15;780(1):73-82. [PubMed:12383482 ]
- Kitaura J, Miki Y, Kato H, Sakakihara Y, Yanagisawa M: Hyperinsulinaemic hypoglycaemia associated with persistent hyperammonaemia. Eur J Pediatr. 1999 May;158(5):410-3. [PubMed:10333126 ]
- Kelley RI: Octenylsuccinic aciduria in children fed protein-hydrolysate formulas containing modified cornstarch. Pediatr Res. 1991 Dec;30(6):564-9. [PubMed:1805153 ]
- Garcia A, Barbas C, Aguilar R, Castro M: Capillary electrophoresis for rapid profiling of organic acidurias. Clin Chem. 1998 Sep;44(9):1905-11. [PubMed:9732975 ]
- Wu N, Yang M, Gaur U, Xu H, Yao Y, Li D: Alpha-Ketoglutarate: Physiological Functions and Applications. Biomol Ther (Seoul). 2016 Jan;24(1):1-8. doi: 10.4062/biomolther.2015.078. Epub 2016 Jan 1. [PubMed:26759695 ]
- Chin RM, Fu X, Pai MY, Vergnes L, Hwang H, Deng G, Diep S, Lomenick B, Meli VS, Monsalve GC, Hu E, Whelan SA, Wang JX, Jung G, Solis GM, Fazlollahi F, Kaweeteerawat C, Quach A, Nili M, Krall AS, Godwin HA, Chang HR, Faull KF, Guo F, Jiang M, Trauger SA, Saghatelian A, Braas D, Christofk HR, Clarke CF, Teitell MA, Petrascheck M, Reue K, Jung ME, Frand AR, Huang J: The metabolite alpha-ketoglutarate extends lifespan by inhibiting ATP synthase and TOR. Nature. 2014 Jun 19;510(7505):397-401. doi: 10.1038/nature13264. Epub 2014 May 14. [PubMed:24828042 ]
- Klysz D, Tai X, Robert PA, Craveiro M, Cretenet G, Oburoglu L, Mongellaz C, Floess S, Fritz V, Matias MI, Yong C, Surh N, Marie JC, Huehn J, Zimmermann V, Kinet S, Dardalhon V, Taylor N: Glutamine-dependent alpha-ketoglutarate production regulates the balance between T helper 1 cell and regulatory T cell generation. Sci Signal. 2015 Sep 29;8(396):ra97. doi: 10.1126/scisignal.aab2610. [PubMed:26420908 ]
- De Vivo DC: The expanding clinical spectrum of mitochondrial diseases. Brain Dev. 1993 Jan-Feb;15(1):1-22. [PubMed:8338207 ]
- Wendisch VF: Microbial Production of Amino Acid-Related Compounds. Adv Biochem Eng Biotechnol. 2017;159:255-269. doi: 10.1007/10_2016_34. [PubMed:27872963 ]
- Elshenawy S, Pinney SE, Stuart T, Doulias PT, Zura G, Parry S, Elovitz MA, Bennett MJ, Bansal A, Strauss JF 3rd, Ischiropoulos H, Simmons RA: The Metabolomic Signature of the Placenta in Spontaneous Preterm Birth. Int J Mol Sci. 2020 Feb 4;21(3). pii: ijms21031043. doi: 10.3390/ijms21031043. [PubMed:32033212 ]
| Show more...
|---|