| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected and Quantified |
|---|
| Creation Date | 2005-11-16 15:48:42 UTC |
|---|
| Update Date | 2023-07-07 20:53:57 UTC |
|---|
| HMDB ID | HMDB0000177 |
|---|
| Secondary Accession Numbers | - HMDB0003412
- HMDB00177
- HMDB03412
|
|---|
| Metabolite Identification |
|---|
| Common Name | Histidine |
|---|
| Description | Histidine (His), also known as L-histidine, is an alpha-amino acid. These are amino acids in which the amino group is attached to the carbon atom immediately adjacent to the carboxylate group (alpha carbon). Amino acids are organic compounds that contain amino (-NH2) and carboxyl (-COOH) functional groups, along with a side chain (R group) specific to each amino acid. Histidine is one of 20 proteinogenic amino acids, i.e., the amino acids used in the biosynthesis of proteins. Histidine is found in all organisms ranging from bacteria to plants to animals. It is classified as an aliphatic, positively charged or basic amino acid. Histidine is a unique amino acid with an imidazole functional group. The acid-base properties of the imidazole side chain are relevant to the catalytic mechanism of many enzymes such as proteases. In catalytic triads, the basic nitrogen of histidine abstracts a proton from serine, threonine, or cysteine to activate it as a nucleophile. In a histidine proton shuttle, histidine is used to quickly shuttle protons. It can do this by abstracting a proton with its basic nitrogen to make a positively charged intermediate and then use another molecule to extract the proton from its acidic nitrogen. Histidine forms complexes with many metal ions. The imidazole sidechain of the histidine residue commonly serves as a ligand in metalloproteins. Histidine was first isolated by German physician Albrecht Kossel in 1896. Histidine is an essential amino acid in humans and other mammals. It was initially thought that it was only essential for infants, but longer-term studies established that it is also essential for adults. Infants four to six months old require 33 mg/kg of histidine. It is not clear how adults make small amounts of histidine, and dietary sources probably account for most of the histidine in the body. Histidine is a precursor for histamine and carnosine biosynthesis. Inborn errors of histidine metabolism, including histidinemia, maple syrup urine disease, propionic acidemia, and tyrosinemia I, exist and are marked by increased histidine levels in the blood. Elevated blood histidine is accompanied by a wide range of symptoms, from mental and physical retardation to poor intellectual functioning, emotional instability, tremor, ataxia and psychosis. Histidine and other imidazole compounds have anti-oxidant, anti-inflammatory and anti-secretory properties (PMID: 9605177 ). The efficacy of L-histidine in protecting inflamed tissue is attributed to the capacity of the imidazole ring to scavenge reactive oxygen species (ROS) generated by cells during acute inflammatory response (PMID: 9605177 ). Histidine, when administered in therapeutic quantities is able to inhibit cytokines and growth factors involved in cell and tissue damage (US patent 6150392). Histidine in medical therapies has its most promising trials in rheumatoid arthritis where up to 4.5 g daily have been used effectively in severely affected patients. Arthritis patients have been found to have low serum histidine levels, apparently because of very rapid removal of histidine from their blood (PMID: 1079527 ). Other patients besides arthritis patients that have been found to be low in serum histidine are those with chronic renal failure. Urinary levels of histidine are reduced in pediatric patients with pneumonia (PMID: 2084459 ). Asthma patients exhibit increased serum levels of histidine over normal controls (PMID: 23517038 ). Serum histidine levels are lower and are negatively associated with inflammation and oxidative stress in obese women (PMID: 23361591 ). Histidine supplementation has been shown to reduce insulin resistance, reduce BMI and fat mass and suppress inflammation and oxidative stress in obese women with metabolic syndrome. Histidine appears to suppress pro-inflammatory cytokine expression, possibly via the NF-κB pathway, in adipocytes (PMID: 23361591 ). Low plasma concentrations of histidine are associated with protein-energy wasting, inflammation, oxidative stress, and greater mortality in chronic kidney disease patients (PMID: 18541578 ). Histidine may have many other possible functions because it is the precursor of the ubiquitous neurohormone-neurotransmitter histamine. Histidine increases histamine in the blood and probably in the brain. Low blood histamine with low serum histidine occurs in rheumatoid arthritis patients. Low blood histamine also occurs in some manic, schizophrenic, high copper and hyperactive groups of psychiatric patients. Histidine is a useful therapy in all patients with low histamine levels (http://www.dcnutrition.com). | Read more...
|---|
| Structure | N[C@@H](CC1=CNC=N1)C(O)=O InChI=1S/C6H9N3O2/c7-5(6(10)11)1-4-2-8-3-9-4/h2-3,5H,1,7H2,(H,8,9)(H,10,11)/t5-/m0/s1 |
|---|
| Synonyms | | Value | Source |
|---|
| (S)-4-(2-Amino-2-carboxyethyl)imidazole | ChEBI | | (S)-alpha-Amino-1H-imidazole-4-propanoic acid | ChEBI | | (S)-alpha-Amino-1H-imidazole-4-propionic acid | ChEBI | | H | ChEBI | | His | ChEBI | | HISTIDINE | ChEBI | | L-(-)-Histidine | ChEBI | | L-Histidin | ChEBI | | (S)-a-Amino-1H-imidazole-4-propanoate | Generator | | (S)-a-Amino-1H-imidazole-4-propanoic acid | Generator | | (S)-alpha-Amino-1H-imidazole-4-propanoate | Generator | | (S)-Α-amino-1H-imidazole-4-propanoate | Generator | | (S)-Α-amino-1H-imidazole-4-propanoic acid | Generator | | (S)-a-Amino-1H-imidazole-4-propionate | Generator | | (S)-a-Amino-1H-imidazole-4-propionic acid | Generator | | (S)-alpha-Amino-1H-imidazole-4-propionate | Generator | | (S)-Α-amino-1H-imidazole-4-propionate | Generator | | (S)-Α-amino-1H-imidazole-4-propionic acid | Generator | | (S)-1H-Imidazole-4-alanine | HMDB | | (S)-2-Amino-3-(4-imidazolyl)propionsaeure | HMDB | | (S)-Histidine | HMDB | | (S)1H-Imidazole-4-alanine | HMDB | | 3-(1H-Imidazol-4-yl)-L-alanine | HMDB | | Amino-1H-imidazole-4-propanoate | HMDB | | Amino-1H-imidazole-4-propanoic acid | HMDB | | Amino-4-imidazoleproprionate | HMDB | | Amino-4-imidazoleproprionic acid | HMDB | | Glyoxaline-5-alanine | HMDB | | Histidine, L isomer | HMDB | | Histidine, L-isomer | HMDB | | L-Isomer histidine | HMDB | | L-Histidine | KEGG |
| Show more...
|---|
| Chemical Formula | C6H9N3O2 |
|---|
| Average Molecular Weight | 155.1546 |
|---|
| Monoisotopic Molecular Weight | 155.069476547 |
|---|
| IUPAC Name | (2S)-2-amino-3-(1H-imidazol-5-yl)propanoic acid |
|---|
| Traditional Name | L-histidine |
|---|
| CAS Registry Number | 71-00-1 |
|---|
| SMILES | N[C@@H](CC1=CNC=N1)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C6H9N3O2/c7-5(6(10)11)1-4-2-8-3-9-4/h2-3,5H,1,7H2,(H,8,9)(H,10,11)/t5-/m0/s1 |
|---|
| InChI Key | HNDVDQJCIGZPNO-YFKPBYRVSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as histidine and derivatives. Histidine and derivatives are compounds containing cysteine or a derivative thereof resulting from reaction of cysteine at the amino group or the carboxy group, or from the replacement of any hydrogen of glycine by a heteroatom. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Carboxylic acids and derivatives |
|---|
| Sub Class | Amino acids, peptides, and analogues |
|---|
| Direct Parent | Histidine and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Histidine or derivatives
- Alpha-amino acid
- L-alpha-amino acid
- Imidazolyl carboxylic acid derivative
- Aralkylamine
- Azole
- Imidazole
- Heteroaromatic compound
- Amino acid
- Carboxylic acid
- Azacycle
- Organoheterocyclic compound
- Monocarboxylic acid or derivatives
- Organic nitrogen compound
- Organooxygen compound
- Organonitrogen compound
- Primary amine
- Primary aliphatic amine
- Hydrocarbon derivative
- Organic oxide
- Organopnictogen compound
- Carbonyl group
- Organic oxygen compound
- Amine
- Aromatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aromatic heteromonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Physiological effect | |
|---|
| Disposition | Biological locationSourceExogenous- Exogenous (HMDB: HMDB0000177)
FoodAnimal originHerb and spiceVegetableFruitNutCereal and cereal productPulseGourdCoffee and coffee productSoyTeaDishBaking goodEggBeverageAquatic originConfectioneryMilk and milk productOther milk productFermented milkFermented milk productUnfermented milk- Milk (Other mammals) (FooDB: FOOD00690)
- Milk (Human) (FooDB: FOOD00666)
- Milk (Cow) (FooDB: FOOD00618)
- Cow milk, pasteurized, vitamin A + D added, 0% fat (FooDB: FOOD00889)
- Cow milk, pasteurized, vitamin A + D added, 1% fat (FooDB: FOOD00890)
- Cow milk, pasteurized, vitamin A + D added, 2% fat (FooDB: FOOD00891)
- Cow milk, pasteurized, vitamin D added, 3.25% fat (FooDB: FOOD00892)
Fat and oilCocoa and cocoa productBaby foodUnclassified food or beverageSnack
|
|---|
| Process | |
|---|
| Role | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | 287 °C | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | 45.6 mg/mL | Not Available | | LogP | -3.32 | CHMELIK,J ET AL. (1991) |
|
|---|
| Experimental Chromatographic Properties | Experimental Collision Cross Sections |
|---|
| Predicted Molecular Properties | | Show more...
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Kovats Retention IndicesUnderivatizedDerivatized| Derivative Name / Structure | SMILES | Kovats RI Value | Column Type | Reference |
|---|
| L-Histidine,1TMS,isomer #1 | C[Si](C)(C)OC(=O)[C@@H](N)CC1=CN=C[NH]1 | 1694.1 | Semi standard non polar | 33892256 | | L-Histidine,1TMS,isomer #2 | C[Si](C)(C)N[C@@H](CC1=CN=C[NH]1)C(=O)O | 1804.7 | Semi standard non polar | 33892256 | | L-Histidine,1TMS,isomer #3 | C[Si](C)(C)N1C=NC=C1C[C@H](N)C(=O)O | 1859.3 | Semi standard non polar | 33892256 | | L-Histidine,2TMS,isomer #1 | C[Si](C)(C)N[C@@H](CC1=CN=C[NH]1)C(=O)O[Si](C)(C)C | 1778.3 | Semi standard non polar | 33892256 | | L-Histidine,2TMS,isomer #1 | C[Si](C)(C)N[C@@H](CC1=CN=C[NH]1)C(=O)O[Si](C)(C)C | 1836.6 | Standard non polar | 33892256 | | L-Histidine,2TMS,isomer #1 | C[Si](C)(C)N[C@@H](CC1=CN=C[NH]1)C(=O)O[Si](C)(C)C | 2385.1 | Standard polar | 33892256 | | L-Histidine,2TMS,isomer #2 | C[Si](C)(C)OC(=O)[C@@H](N)CC1=CN=CN1[Si](C)(C)C | 1856.5 | Semi standard non polar | 33892256 | | L-Histidine,2TMS,isomer #2 | C[Si](C)(C)OC(=O)[C@@H](N)CC1=CN=CN1[Si](C)(C)C | 1801.7 | Standard non polar | 33892256 | | L-Histidine,2TMS,isomer #2 | C[Si](C)(C)OC(=O)[C@@H](N)CC1=CN=CN1[Si](C)(C)C | 2504.8 | Standard polar | 33892256 | | L-Histidine,2TMS,isomer #3 | C[Si](C)(C)N([C@@H](CC1=CN=C[NH]1)C(=O)O)[Si](C)(C)C | 1911.9 | Semi standard non polar | 33892256 | | L-Histidine,2TMS,isomer #3 | C[Si](C)(C)N([C@@H](CC1=CN=C[NH]1)C(=O)O)[Si](C)(C)C | 1927.2 | Standard non polar | 33892256 | | L-Histidine,2TMS,isomer #3 | C[Si](C)(C)N([C@@H](CC1=CN=C[NH]1)C(=O)O)[Si](C)(C)C | 2536.1 | Standard polar | 33892256 | | L-Histidine,2TMS,isomer #4 | C[Si](C)(C)N[C@@H](CC1=CN=CN1[Si](C)(C)C)C(=O)O | 1918.3 | Semi standard non polar | 33892256 | | L-Histidine,2TMS,isomer #4 | C[Si](C)(C)N[C@@H](CC1=CN=CN1[Si](C)(C)C)C(=O)O | 1827.6 | Standard non polar | 33892256 | | L-Histidine,2TMS,isomer #4 | C[Si](C)(C)N[C@@H](CC1=CN=CN1[Si](C)(C)C)C(=O)O | 2501.0 | Standard polar | 33892256 | | L-Histidine,3TMS,isomer #1 | C[Si](C)(C)OC(=O)[C@H](CC1=CN=C[NH]1)N([Si](C)(C)C)[Si](C)(C)C | 1938.6 | Semi standard non polar | 33892256 | | L-Histidine,3TMS,isomer #1 | C[Si](C)(C)OC(=O)[C@H](CC1=CN=C[NH]1)N([Si](C)(C)C)[Si](C)(C)C | 1959.4 | Standard non polar | 33892256 | | L-Histidine,3TMS,isomer #1 | C[Si](C)(C)OC(=O)[C@H](CC1=CN=C[NH]1)N([Si](C)(C)C)[Si](C)(C)C | 2231.5 | Standard polar | 33892256 | | L-Histidine,3TMS,isomer #2 | C[Si](C)(C)N[C@@H](CC1=CN=CN1[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1941.1 | Semi standard non polar | 33892256 | | L-Histidine,3TMS,isomer #2 | C[Si](C)(C)N[C@@H](CC1=CN=CN1[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1856.2 | Standard non polar | 33892256 | | L-Histidine,3TMS,isomer #2 | C[Si](C)(C)N[C@@H](CC1=CN=CN1[Si](C)(C)C)C(=O)O[Si](C)(C)C | 2187.4 | Standard polar | 33892256 | | L-Histidine,3TMS,isomer #3 | C[Si](C)(C)N([C@@H](CC1=CN=CN1[Si](C)(C)C)C(=O)O)[Si](C)(C)C | 2065.9 | Semi standard non polar | 33892256 | | L-Histidine,3TMS,isomer #3 | C[Si](C)(C)N([C@@H](CC1=CN=CN1[Si](C)(C)C)C(=O)O)[Si](C)(C)C | 1975.4 | Standard non polar | 33892256 | | L-Histidine,3TMS,isomer #3 | C[Si](C)(C)N([C@@H](CC1=CN=CN1[Si](C)(C)C)C(=O)O)[Si](C)(C)C | 2310.6 | Standard polar | 33892256 | | L-Histidine,4TMS,isomer #1 | C[Si](C)(C)OC(=O)[C@H](CC1=CN=CN1[Si](C)(C)C)N([Si](C)(C)C)[Si](C)(C)C | 2129.7 | Semi standard non polar | 33892256 | | L-Histidine,4TMS,isomer #1 | C[Si](C)(C)OC(=O)[C@H](CC1=CN=CN1[Si](C)(C)C)N([Si](C)(C)C)[Si](C)(C)C | 2011.6 | Standard non polar | 33892256 | | L-Histidine,4TMS,isomer #1 | C[Si](C)(C)OC(=O)[C@H](CC1=CN=CN1[Si](C)(C)C)N([Si](C)(C)C)[Si](C)(C)C | 2116.0 | Standard polar | 33892256 | | L-Histidine,1TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)[C@@H](N)CC1=CN=C[NH]1 | 1926.3 | Semi standard non polar | 33892256 | | L-Histidine,1TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)N[C@@H](CC1=CN=C[NH]1)C(=O)O | 2025.1 | Semi standard non polar | 33892256 | | L-Histidine,1TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)N1C=NC=C1C[C@H](N)C(=O)O | 2140.1 | Semi standard non polar | 33892256 | | L-Histidine,2TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)N[C@@H](CC1=CN=C[NH]1)C(=O)O[Si](C)(C)C(C)(C)C | 2219.9 | Semi standard non polar | 33892256 | | L-Histidine,2TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)N[C@@H](CC1=CN=C[NH]1)C(=O)O[Si](C)(C)C(C)(C)C | 2293.0 | Standard non polar | 33892256 | | L-Histidine,2TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)N[C@@H](CC1=CN=C[NH]1)C(=O)O[Si](C)(C)C(C)(C)C | 2523.7 | Standard polar | 33892256 | | L-Histidine,2TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC(=O)[C@@H](N)CC1=CN=CN1[Si](C)(C)C(C)(C)C | 2356.2 | Semi standard non polar | 33892256 | | L-Histidine,2TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC(=O)[C@@H](N)CC1=CN=CN1[Si](C)(C)C(C)(C)C | 2258.0 | Standard non polar | 33892256 | | L-Histidine,2TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC(=O)[C@@H](N)CC1=CN=CN1[Si](C)(C)C(C)(C)C | 2593.9 | Standard polar | 33892256 | | L-Histidine,2TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)N([C@@H](CC1=CN=C[NH]1)C(=O)O)[Si](C)(C)C(C)(C)C | 2332.5 | Semi standard non polar | 33892256 | | L-Histidine,2TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)N([C@@H](CC1=CN=C[NH]1)C(=O)O)[Si](C)(C)C(C)(C)C | 2369.5 | Standard non polar | 33892256 | | L-Histidine,2TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)N([C@@H](CC1=CN=C[NH]1)C(=O)O)[Si](C)(C)C(C)(C)C | 2607.4 | Standard polar | 33892256 | | L-Histidine,2TBDMS,isomer #4 | CC(C)(C)[Si](C)(C)N[C@@H](CC1=CN=CN1[Si](C)(C)C(C)(C)C)C(=O)O | 2427.2 | Semi standard non polar | 33892256 | | L-Histidine,2TBDMS,isomer #4 | CC(C)(C)[Si](C)(C)N[C@@H](CC1=CN=CN1[Si](C)(C)C(C)(C)C)C(=O)O | 2276.5 | Standard non polar | 33892256 | | L-Histidine,2TBDMS,isomer #4 | CC(C)(C)[Si](C)(C)N[C@@H](CC1=CN=CN1[Si](C)(C)C(C)(C)C)C(=O)O | 2578.1 | Standard polar | 33892256 | | L-Histidine,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CC1=CN=C[NH]1)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2588.3 | Semi standard non polar | 33892256 | | L-Histidine,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CC1=CN=C[NH]1)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2584.0 | Standard non polar | 33892256 | | L-Histidine,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CC1=CN=C[NH]1)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2487.9 | Standard polar | 33892256 | | L-Histidine,3TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)N[C@@H](CC1=CN=CN1[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2611.9 | Semi standard non polar | 33892256 | | L-Histidine,3TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)N[C@@H](CC1=CN=CN1[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2507.3 | Standard non polar | 33892256 | | L-Histidine,3TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)N[C@@H](CC1=CN=CN1[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2456.0 | Standard polar | 33892256 | | L-Histidine,3TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)N([C@@H](CC1=CN=CN1[Si](C)(C)C(C)(C)C)C(=O)O)[Si](C)(C)C(C)(C)C | 2745.5 | Semi standard non polar | 33892256 | | L-Histidine,3TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)N([C@@H](CC1=CN=CN1[Si](C)(C)C(C)(C)C)C(=O)O)[Si](C)(C)C(C)(C)C | 2591.8 | Standard non polar | 33892256 | | L-Histidine,3TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)N([C@@H](CC1=CN=CN1[Si](C)(C)C(C)(C)C)C(=O)O)[Si](C)(C)C(C)(C)C | 2528.2 | Standard polar | 33892256 | | L-Histidine,4TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CC1=CN=CN1[Si](C)(C)C(C)(C)C)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2961.0 | Semi standard non polar | 33892256 | | L-Histidine,4TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CC1=CN=CN1[Si](C)(C)C(C)(C)C)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2808.9 | Standard non polar | 33892256 | | L-Histidine,4TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CC1=CN=CN1[Si](C)(C)C(C)(C)C)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2484.2 | Standard polar | 33892256 |
| Show more...
|---|
| Spectra |
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental GC-MS | GC-MS Spectrum - Histidine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-0udi-0940000000-43b9b035189af65e19cb | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Histidine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-0udi-0910000000-fc3ecec9d4ff3aa8cb09 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Histidine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (Non-derivatized) | splash10-0udi-0910000000-8284a673a35f7ac2241f | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Histidine GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-0fk9-8910000000-a3231b1a418b5798ecac | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Histidine GC-MS (3 TMS) | splash10-0udi-1920000000-227e8116769731104e82 | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Histidine GC-MS (4 TMS) | splash10-0ufu-2942100000-31605fd50ecc1f5be0ed | 2014-06-16 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Histidine GC-EI-TOF (Non-derivatized) | splash10-0udi-0940000000-43b9b035189af65e19cb | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Histidine GC-EI-TOF (Non-derivatized) | splash10-0udi-0910000000-fc3ecec9d4ff3aa8cb09 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Histidine GC-EI-TOF (Non-derivatized) | splash10-0udi-0910000000-8284a673a35f7ac2241f | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Histidine GC-EI-QQ (Non-derivatized) | splash10-0002-2901100000-ae0f3865934743de764c | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Histidine GC-EI-TOF (Non-derivatized) | splash10-0fk9-8910000000-a3231b1a418b5798ecac | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Histidine GC-MS (Non-derivatized) | splash10-0udi-1920000000-227e8116769731104e82 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Histidine GC-MS (Non-derivatized) | splash10-0ufu-2942100000-31605fd50ecc1f5be0ed | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Histidine GC-EI-TOF (Non-derivatized) | splash10-0ue9-0900000000-75e38a3f347210c301b0 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Histidine GC-EI-TOF (Non-derivatized) | splash10-0udi-0910000000-c54bd92cc112eec7e509 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Histidine GC-MS (Non-derivatized) - 70eV, Positive | splash10-001i-9500000000-1d29684ccb6383687865 | 2016-09-22 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Histidine GC-MS (1 TMS) - 70eV, Positive | splash10-03di-6900000000-1dd400a16b8067f4daea | 2017-10-06 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Histidine GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Histidine GC-MS (TMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Histidine GC-MS (TMS_1_3) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Histidine GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Histidine GC-MS (TBDMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Histidine GC-MS (TBDMS_1_3) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-001i-9100000000-ae0a0b81669663b3b4a5 | 2018-05-25 | Not Available | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine Quattro_QQQ 10V, Positive-QTOF (Annotated) | splash10-0a4i-0900000000-21530fac9975dc1edc7c | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine Quattro_QQQ 25V, Positive-QTOF (Annotated) | splash10-053r-9300000000-a946c375931adb366aae | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine Quattro_QQQ 40V, Positive-QTOF (Annotated) | splash10-053r-9000000000-61a3602d80bc0d54c579 | 2012-07-24 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-0a4i-0900000000-7d63b501889c2628f4f0 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-001i-0900000000-d4ccd4e44ca818b4cb17 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-0a4i-0900000000-811aee5724fe0999c134 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-0a4i-2900000000-052753eb699ae7cb4cb9 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-0a4i-0900000000-3e145a05e6a71a0a2f56 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-0a4i-0900000000-143a4b9d88183bc5abe7 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-03di-0900000000-a8c213472da676a241f2 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive-QTOF | splash10-001i-0900000000-dfdf9562f155abb5fdeb | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-0uk9-0879331100-8434b5b9e7e5700c2353 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-000i-0900000000-aaaaadcecd4cd7c94e1e | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-0udi-0900000000-90c4cbd09d7abc003013 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-0udi-0900000000-1381708d18c24486773d | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-0w29-0696321100-eb62fc608270d1151707 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-000i-0900000000-3450566ff38e6506a70f | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-01q9-7900000000-124509a66476629ce10a | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative-QTOF | splash10-0udi-0900000000-efa6a21b1be16fda0802 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-QQ (API3000, Applied Biosystems) 10V, Negative-QTOF | splash10-0udi-0900000000-137840e69da48c6114e4 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-QQ (API3000, Applied Biosystems) 20V, Negative-QTOF | splash10-0udu-3900000000-06d50e90c797793a72d7 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-QQ (API3000, Applied Biosystems) 30V, Negative-QTOF | splash10-0006-9100000000-de7c9eb8068ef4f39f85 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-QQ (API3000, Applied Biosystems) 40V, Negative-QTOF | splash10-00l6-9000000000-ebf778679d4c6d6a6057 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-QQ (API3000, Applied Biosystems) 50V, Negative-QTOF | splash10-014i-9000000000-08d0188684283d1b2372 | 2012-08-31 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Histidine LC-ESI-QQ (API3000, Applied Biosystems) 10V, Positive-QTOF | splash10-0a4i-0900000000-46c2f608d27f0381a039 | 2012-08-31 | HMDB team, MONA | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Experimental 1D NMR | 1H NMR Spectrum (1D, 600 MHz, H2O, experimental) | 2021-10-10 | Wishart Lab | View Spectrum | | Experimental 1D NMR | 13C NMR Spectrum (1D, 400 MHz, H2O, experimental) | 2021-10-10 | Wishart Lab | View Spectrum | | Experimental 1D NMR | 1H NMR Spectrum (1D, 600 MHz, H2O, experimental) | 2021-10-10 | Wishart Lab | View Spectrum | | Experimental 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, H2O, experimental) | 2012-12-05 | Wishart Lab | View Spectrum |
IR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M-H]-) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+H]+) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+Na]+) | 2023-02-03 | FELIX lab | View Spectrum |
| Show more...
|---|
| Biological Properties |
|---|
| Cellular Locations | |
|---|
| Biospecimen Locations | - Blood
- Breast Milk
- Cerebrospinal Fluid (CSF)
- Feces
- Saliva
- Sweat
- Urine
|
|---|
| Tissue Locations | - All Tissues
- Placenta
- Prostate
|
|---|
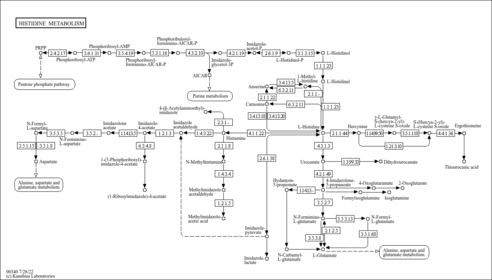
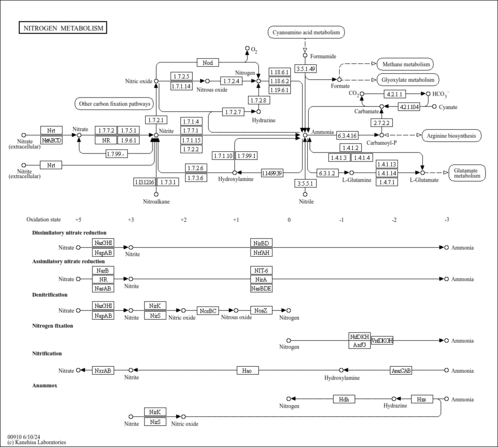
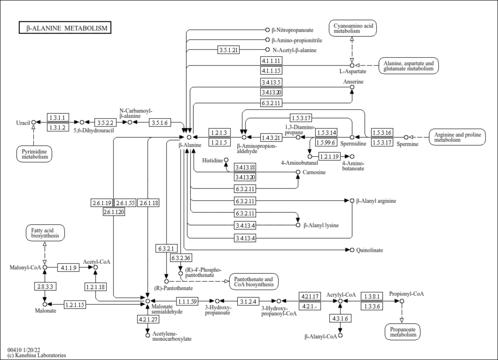
| Pathways | |
|---|
| Normal Concentrations |
|---|
| |
| Blood | Detected and Quantified | 75.0 +/- 20.0 uM | Newborn (0-30 days old) | Not Specified | Normal | | details | | Blood | Detected and Quantified | 86.0 +/- 10.0 uM | Children (1-13 years old) | Male | Normal | | details | | Blood | Detected and Quantified | 89.0 +/- 11.0 uM | Adult (>18 years old) | Male | Normal | | details | | Blood | Detected and Quantified | 83.0 +/- 14.0 uM | Adult (>18 years old) | Female | Normal | | details | | Blood | Detected and Quantified | 143.1 +/- 27.3 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 32-107 uM | Newborn (0-30 days old) | Both | Normal | | details | | Blood | Detected and Quantified | 20-83 uM | Infant (1 - 3 months old) | Both | Normal | | details | | Blood | Detected and Quantified | 22-108 uM | Children (3 months - 6 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 35-106 uM | Children (6 - 18 years old) | Both | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 124 (104-150) uM | Newborn (0-30 days old) | Not Available | Normal | | details | | Blood | Detected and Quantified | 90 +/- 9 uM | Adult (>18 years old) | Male | Normal | | details | | Blood | Detected and Quantified | 86 +/- 9 uM | Adult (>18 years old) | Female | Normal | | details | | Blood | Detected and Quantified | 106 (91.7-124) uM | Infant (0-1 year old) | Not Available | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 32.226 uM | Children (1 - 13 years old) | Not Specified | Normal | | details | | Blood | Detected and Quantified | 90.233 uM | Children (1 - 13 years old) | Not Specified | Normal | | details | | Blood | Detected and Quantified | 128.904 uM | Children (1-13 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 225.581 uM | Children (1 - 13 years old) | Not Specified | Normal | | details | | Blood | Detected and Quantified | 73.51(11.42) uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 122.459 +/- 12.890 uM | Children (1 - 13 years old) | Male | Normal | | details | | Blood | Detected and Quantified | 128.904-354.485 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Blood | Detected and Quantified | 225.581-386.711 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Blood | Detected and Quantified | 30-138 uM | Infant (0-1 year old) | Not Specified | Normal | | details | | Blood | Detected and Quantified | 83 +/- 14 uM | Adult (>18 years old) | Female | Normal | | details | | Blood | Detected and Quantified | 40.00-120.0 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 50 - 110 uM | Children (1 - 13 years old) | Not Specified | Normal | | details | | Blood | Detected and Quantified | 103.13-141.80 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Blood | Detected and Quantified | 89.3 +/- 10.8 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 77.342-238.472 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Blood | Detected and Quantified | 96.678-225.581 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Blood | Detected and Quantified | 89 +/- 11 uM | Adult (>18 years old) | Male | Normal | | details | | Blood | Detected and Quantified | 83.4(70.1-97.4) uM | Children (1-13 uears old) | Both | Normal | | details | | Blood | Detected and Quantified | 82.0 (72.0-92.0) uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 84 +/- 14 uM | Children (1 - 13 years old) | Not Specified | Normal | | details | | Blood | Detected and Quantified | 131.2 +/- 37.3 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Children (1-13 years old) | Male | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Children (1-13 years old) | Male | Normal | | details | | Breast Milk | Detected and Quantified | 20.3 +/- 6.9 uM | Adult (>18 years old) | Female | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 7.0897-19.980 uM | Children (1 - 13 years old) | Male | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 14.0 +/- 8.0 uM | Not Specified | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 12.4 +/- 3.5 uM | Children (1 - 13 years old) | Female | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 11.6 +/- 4.4 uM | Children (1 - 13 years old) | Male | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 3-18 uM | Newborn (0-30 days old) | Not Specified | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 20.7 +/- 7.5 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 16.8 +/- 2.7 uM | Adult (>18 years old) | Male | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 18.6 +/- 3.3 uM | Adult (>18 years old) | Female | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 16.4 +/- 6.1 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 10.9 (10.4-11.4) uM | Adult (>18 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 0-17 uM | Children (0 - 10 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 4.6-23 uM | Adolescent (>11 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 11.9 +/- 1.7 uM | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Children (1-13 years old) | Both | Normal | | details | | Feces | Detected and Quantified | 1718.716 (1310.521-2126.911) nmol/g wet feces | Infant (0-1 year old) | Not Specified | Normal | | details | | Feces | Detected and Quantified | 2298.782 (1224.585-3372.980) nmol/g wet feces | Infant (0-1 year old) | Not Specified | Normal | | details | | Feces | Detected and Quantified | 966.778 (451.163-1482.392) nmol/g wet feces | Infant (0-1 year old) | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Children (1-13 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | Normal | | details | | Feces | Detected and Quantified | 60 +/- 50 nmol/g wet feces | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Children (1-13 years old) | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Children (1-13 years old) | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected and Quantified | 24.99 +/- 12.94 uM | Adult (>18 years old) | Female | Normal | | details | | Saliva | Detected and Quantified | 6.02 +/- 4.39 uM | Adult (>18 years old) | Female | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected and Quantified | 9.93 +/- 9.50 uM | Adult (>18 years old) | Female | Normal | | details | | Saliva | Detected and Quantified | <0.25 uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | <0.25 uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 9.4 +/- 4.9 uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | 18.67 +/- 10.20 uM | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected and Quantified | 8.2 +/- 1.2 uM | Adult (>18 years old) | Male | Normal | | details | | Saliva | Detected and Quantified | >10 uM | Adult (>18 years old) | Both | Normal | | details | | Sweat | Detected and Quantified | 170 uM | Adult (60 years old) | Male | Normal | | details | | Sweat | Detected and Quantified | < 10 uM | Adult (40 years old) | Male | Normal | | details | | Sweat | Detected but not Quantified | Not Quantified | Adult | Both | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Children (1-13 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 45.22 - 167.55 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 40.70 - 194.35 umol/mmol creatinine | Infant (1 - 6 months old) | Both | Normal | | details | | Urine | Detected and Quantified | 52.01 - 157.83 umol/mmol creatinine | Infant (6 months - <1 year old) | Both | Normal | | details | | Urine | Detected and Quantified | 49.18 - 163.03 umol/mmol creatinine | Children (1 - 2 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 38.44 - 144.83 umol/mmol creatinine | Children (2 - 4 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 24.19 - 122.67 umol/mmol creatinine | Children (4 - 13 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 14.47 - 86.72 umol/mmol creatinine | Adolescent (13 - 21 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 14.47 - 86.72 umol/mmol creatinine | Adult (>21 years old) | Both | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 15.16-224.31 umol/mmol creatinine | Adolescent (13-18 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 101.9 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 25.79-212.81 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 45-100 umol/mmol creatinine | Adult (>18 years old) | Female | Normal | | details | | Urine | Detected and Quantified | 9.16-127.60 umol/mmol creatinine | Adult (>18 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 87.2 (23.0-151.0) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 16.4-433.51 umol/mmol creatinine | Infant (0-1 year old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 47.12 umol/mmol creatinine | Adult (>18 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 0.0036 - 0.0376 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 19.07-342.31 umol/mmol creatinine | Newborn (0-30 days old) | Female | Normal | | details | | Urine | Detected and Quantified | 4.78-199.77 umol/mmol creatinine | Newborn (0-30 days old) | Male | Normal | | details | | Urine | Detected and Quantified | 18.0667 +/- 9.133 umol/mmol creatinine | Children (1 - 13 years old) | Female | Normal | | details | | Urine | Detected and Quantified | 119.45 +/- 87.37 umol/mmol creatinine | Newborn (0-30 days old) | Female | Normal | | details | | Urine | Detected and Quantified | 31.667 +/- 22.800 umol/mmol creatinine | Children (1 - 13 years old) | Female | Normal | | details | | Urine | Detected and Quantified | 43 (17-90) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 150.04 +/- 94 umol/mmol creatinine | Children (0-2 years old) | Female | Normal | | details | | Urine | Detected and Quantified | 154.26 +/- 82.28 umol/mmol creatinine | Children (0-2 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 60.14 +/- 47.11 umol/mmol creatinine | Newborn (0-30 days old) | Male | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 31.200 +/- 15.133 umol/mmol creatinine | Children (1 - 13 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 46.933 +/- 30 umol/mmol creatinine | Children (1 - 13 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 12.6 +/- 3.5 umol/mmol creatinine | Infant (1 - 6 months old) | Both | Normal | | details | | Urine | Detected and Quantified | 41.267-256.307 umol/mmol creatinine | Infant (0-1 year old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 15.0 +/- 3.7 umol/mmol creatinine | Infant (6 months - <1 year old) | Both | Normal | | details | | Urine | Detected and Quantified | 8.3 +/- 2.1 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 48.30-384.37 umol/mmol creatinine | Children (1-13 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 154.67 (80-229.33) umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | - Geigy Scientific ...
- West Cadwell, N.J...
- Basel, Switzerlan...
| details | | Urine | Detected and Quantified | 31.0 +/- 15.0 umol/mmol creatinine | Children (1-13 years old) | Male | Normal | - Geigy Scientific ...
- West Cadwell, N.J...
- Basel, Switzerlan...
| details | | Urine | Detected and Quantified | 85.0 +/- 33.0 umol/mmol creatinine | Adult (>18 years old) | Male | Normal | - Geigy Scientific ...
- West Cadwell, N.J...
- Basel, Switzerlan...
| details | | Urine | Detected and Quantified | 62.0 +/- 28.0 umol/mmol creatinine | Adult (>18 years old) | Female | Normal | - Geigy Scientific ...
- West Cadwell, N.J...
- Basel, Switzerlan...
| details | | Urine | Detected and Quantified | 60.75 +/- 27.45 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 45.6 (21.7-71.5) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 66-106 umol/mmol creatinine | Adult (>18 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 26.017-298.066 umol/mmol creatinine | Children (1-13 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 88 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 190.3063 +/- 67.9627 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 194.90 +/- 144.31 umol/mmol creatinine | Infant (0-1 year old) | Both | Normal | | details | | Urine | Detected and Quantified | 7.914 +/- 6.784 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 9.05 +/- 6.78 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 10.18 +/- 7.91 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 63.73 (36.37-98.87) umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 237.43 +/- 113.06 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 316.57 +/- 146.98 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 463.55 +/- 192.20 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 62.585 +/- 67.649 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Normal | | details | | Urine | Detected and Quantified | 21.09-188.1 umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 53.50(29.22-137.2) umol/mmol creatinine | Newborn (0-30 days old) | Female | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Children (1-13 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 60.27(18.73-193.6) umol/mmol creatinine | Newborn (0-30 days old) | Male | Normal | | details | | Urine | Detected and Quantified | 30.32-242.86 umol/mmol creatinine | Children (1-13 years old) | Not Specified | Normal | | details |
| Show more...
|---|
| Abnormal Concentrations |
|---|
| |
| Blood | Detected and Quantified | 86.0 +/- 21.4 uM | Children (1-13 years old) | Both | Obesity | | details | | Blood | Detected and Quantified | 90.7 +/- 18.4 uM | Children (1-13 years old) | Both | Obesity | | details | | Blood | Detected and Quantified | 46.1 +/- 17.5 uM | Adult (>18 years old) | Both | Heart Transplant | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Blood | Detected and Quantified | 84 +/- 11 uM | Adult (>18 years old) | Male | Schizophrenia | | details | | Blood | Detected and Quantified | 84 +/- 14 uM | Adult (>18 years old) | Female | Schizophrenia | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Schizophrenia | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Pancreatic cancer | | details | | Blood | Detected and Quantified | 130 uM | Infant (0-1 year old) | Male | Fumaric aciduria | | details | | Blood | Detected and Quantified | 80.88(11.04) uM | Adult (>18 years old) | Both | Heart failure with preserved ejection fraction | | details | | Blood | Detected and Quantified | 509.17 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Blood | Detected and Quantified | 92.6(75.9-112.4) uM | Children (1-13 uears old) | Both | Environmental enteric dysfunction | | details | | Blood | Detected and Quantified | 1108.3 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Blood | Detected and Quantified | 257.807-483.389 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Blood | Detected and Quantified | 386.711 +/- 128.904 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Blood | Detected and Quantified | 116.68 +/- 18.15 uM | Elderly (>65 years old) | Both | Alzheimer's disease | | details | | Blood | Detected and Quantified | 412.492-1115.0169 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Blood | Detected and Quantified | 412.492-837.874 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Blood | Detected and Quantified | 470.498-928.107 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Blood | Detected and Quantified | 483.389-708.970 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Blood | Detected and Quantified | 509.170-999.00357 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Blood | Detected and Quantified | 515.615-1063.455 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Blood | Detected and Quantified | 580.0666-1173.0236 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Blood | Detected and Quantified | 708.970-1501.728 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Blood | Detected and Quantified | 72 +/- 19 uM | Children (1 - 13 years old) | Male | Histidinuria | | details | | Blood | Detected and Quantified | 741.196-947.442 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Blood | Detected and Quantified | 837.874 +/- 290.0333 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Blood | Detected and Quantified | 837.874-1630.632 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Blood | Detected and Quantified | 863.655-1115.0169 uM | Children (1-13 years old) | Both | Histidinemia | | details | | Blood | Detected and Quantified | 67.0 (64.0-70.0) uM | Adult (>18 years old) | Both | Dengue fever | | details | | Blood | Detected and Quantified | 80.5 +/- 4.2 uM | Adult (>18 years old) | Both | Epilepsy | | details | | Blood | Detected and Quantified | 107.0 +/- 6.5 uM | Adult (>18 years old) | Both | Epilepsy | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 58.00666 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 116.0133 uM | Children (1 - 13 years old) | Male | Histidinemia | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 20-29 uM | Newborn (0-30 days old) | Both | Pyridoxamine 5-prime-phosphate oxidase deficiency | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 21.4 +/- 6.6 uM | Children (1-13 years old) | Not Specified | Leukemia | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 10.3 +/- 3.7 uM | Children (1-13 years old) | Not Specified | Leukemia | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 130.0 uM | Adult (>18 years old) | Both | Histidinemia | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 14.4 +/- 6.32 uM | Adult (>18 years old) | Both | Progressive dementia | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal Cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Children (1-13 years old) | Both | Autism | | details | | Feces | Detected but not Quantified | Not Quantified | Children (1-13 years old) | Both | celiac disease | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Both | Crohns disease | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Both | Ulcerative colitis | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Not Specified | Unclassified IBD | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Not Specified | Crohns disease | | details | | Feces | Detected but not Quantified | Not Quantified | Children (6 - 18 years old) | Not Specified | Ulcerative colitis | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal Cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Children (1-13 years old) | Not Specified | celiac disease | | details | | Feces | Detected but not Quantified | Not Quantified | Children (1-13 years old) | Not Specified | Treated celiac disease | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Oral cancer | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | Breast cancer | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Pancreatic cancer | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Periodontal diseases | | details | | Saliva | Detected and Quantified | 16.96 +/- 7.43 uM | Adult (>18 years old) | Male | Alzheimer's disease | | details | | Saliva | Detected and Quantified | 23.31 +/- 8.62 uM | Adult (>18 years old) | Male | Frontotemporal lobe dementia | | details | | Saliva | Detected and Quantified | 17.50 +/- 4.53 uM | Adult (>18 years old) | Both | Lewy body disease | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | Epithelial ovarian cancer | | details | | Urine | Detected and Quantified | 0.0033 - 0.0186 umol/mmol creatinine | Adult (>18 years old) | Both | ADPKD | | details | | Urine | Detected and Quantified | 30.5 +/- 4.3 umol/mmol creatinine | Adult (>18 years old) | Both | Alzheimer's disease | | details | | Urine | Detected and Quantified | 276 umol/mmol creatinine | Children (1 - 13 years old) | Male | Histidinuria | | details | | Urine | Detected and Quantified | 595 umol/mmol creatinine | Children (1 - 13 years old) | Male | Histidinuria | | details | | Urine | Detected and Quantified | 246.700-642.700 umol/mmol creatinine | Infant (0-1 year old) | Not Specified | Fumaric aciduria | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Urine | Detected and Quantified | 145.1681 +/- 74.4203 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Eosinophilic esophagitis | | details | | Urine | Detected and Quantified | 103.2581 +/- 61.6795 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Gastroesophageal reflux disease | | details | | Urine | Detected and Quantified | 105.3 +/- 12.7 umol/mmol creatinine | Adult (>18 years old) | Both | 3-Hydroxy-3- methylglutaryl-CoA lyase (HL) deficency | | details | | Urine | Detected and Quantified | 65.5 +/- 11.3 umol/mmol creatinine | Adult (>18 years old) | Both | 3-Hydroxy-3- methylglutaryl-CoA lyase (HL) deficency | | details | | Urine | Detected and Quantified | 52.9 +/- 12.5 umol/mmol creatinine | Adult (>18 years old) | Both | Argininosuccinic aciduria (ASL) | | details | | Urine | Detected and Quantified | 154.3 +/- 13.7 umol/mmol creatinine | Adult (>18 years old) | Both | Propionic acidemia | | details | | Urine | Detected and Quantified | 52 +/- 29 umol/mmol creatinine | Adult (>18 years old) | Both | Tyrosinemia I | | details | | Urine | Detected and Quantified | 37.9 +/- 9.8 umol/mmol creatinine | Adult (>18 years old) | Both | Tyrosinemia I | | details | | Urine | Detected and Quantified | 62.6 +/- 12.3 umol/mmol creatinine | Adult (>18 years old) | Both | Phenylketonuria | | details | | Urine | Detected and Quantified | 68 +/- 19.9 umol/mmol creatinine | Adult (>18 years old) | Both | Phenylketonuria | | details | | Urine | Detected and Quantified | 171.8 +/- 4.3 umol/mmol creatinine | Adult (>18 years old) | Both | Maple syrup urine disease | | details | | Urine | Detected and Quantified | 180.1 +/- 12.4 umol/mmol creatinine | Adult (>18 years old) | Both | Aminoaciduria | | details | | Urine | Detected and Quantified | 64.206 +/- 58.48 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Eosinophilic esophagitis | | details |
| Show more...
|---|
| Associated Disorders and Diseases |
|---|
| Disease References | | Dengue fever |
|---|
- Klassen P, Furst P, Schulz C, Mazariegos M, Solomons NW: Plasma free amino acid concentrations in healthy Guatemalan adults and in patients with classic dengue. Am J Clin Nutr. 2001 Mar;73(3):647-52. [PubMed:11237944 ]
| | Epilepsy |
|---|
- Rainesalo S, Keranen T, Palmio J, Peltola J, Oja SS, Saransaari P: Plasma and cerebrospinal fluid amino acids in epileptic patients. Neurochem Res. 2004 Jan;29(1):319-24. [PubMed:14992292 ]
| | Alzheimer's disease |
|---|
- Fonteh AN, Harrington RJ, Tsai A, Liao P, Harrington MG: Free amino acid and dipeptide changes in the body fluids from Alzheimer's disease subjects. Amino Acids. 2007 Feb;32(2):213-24. Epub 2006 Oct 10. [PubMed:17031479 ]
- Tsuruoka M, Hara J, Hirayama A, Sugimoto M, Soga T, Shankle WR, Tomita M: Capillary electrophoresis-mass spectrometry-based metabolome analysis of serum and saliva from neurodegenerative dementia patients. Electrophoresis. 2013 Oct;34(19):2865-72. doi: 10.1002/elps.201300019. Epub 2013 Sep 6. [PubMed:23857558 ]
| | Colorectal cancer |
|---|
- Ritchie SA, Ahiahonu PW, Jayasinghe D, Heath D, Liu J, Lu Y, Jin W, Kavianpour A, Yamazaki Y, Khan AM, Hossain M, Su-Myat KK, Wood PL, Krenitsky K, Takemasa I, Miyake M, Sekimoto M, Monden M, Matsubara H, Nomura F, Goodenowe DB: Reduced levels of hydroxylated, polyunsaturated ultra long-chain fatty acids in the serum of colorectal cancer patients: implications for early screening and detection. BMC Med. 2010 Feb 15;8:13. doi: 10.1186/1741-7015-8-13. [PubMed:20156336 ]
- Ni Y, Xie G, Jia W: Metabonomics of human colorectal cancer: new approaches for early diagnosis and biomarker discovery. J Proteome Res. 2014 Sep 5;13(9):3857-70. doi: 10.1021/pr500443c. Epub 2014 Aug 14. [PubMed:25105552 ]
- Goedert JJ, Sampson JN, Moore SC, Xiao Q, Xiong X, Hayes RB, Ahn J, Shi J, Sinha R: Fecal metabolomics: assay performance and association with colorectal cancer. Carcinogenesis. 2014 Sep;35(9):2089-96. doi: 10.1093/carcin/bgu131. Epub 2014 Jul 18. [PubMed:25037050 ]
- Brown DG, Rao S, Weir TL, O'Malia J, Bazan M, Brown RJ, Ryan EP: Metabolomics and metabolic pathway networks from human colorectal cancers, adjacent mucosa, and stool. Cancer Metab. 2016 Jun 6;4:11. doi: 10.1186/s40170-016-0151-y. eCollection 2016. [PubMed:27275383 ]
- Sinha R, Ahn J, Sampson JN, Shi J, Yu G, Xiong X, Hayes RB, Goedert JJ: Fecal Microbiota, Fecal Metabolome, and Colorectal Cancer Interrelations. PLoS One. 2016 Mar 25;11(3):e0152126. doi: 10.1371/journal.pone.0152126. eCollection 2016. [PubMed:27015276 ]
| | Schizophrenia |
|---|
- Bjerkenstedt L, Edman G, Hagenfeldt L, Sedvall G, Wiesel FA: Plasma amino acids in relation to cerebrospinal fluid monoamine metabolites in schizophrenic patients and healthy controls. Br J Psychiatry. 1985 Sep;147:276-82. [PubMed:2415198 ]
| | Fumarase deficiency |
|---|
- Allegri G, Fernandes MJ, Scalco FB, Correia P, Simoni RE, Llerena JC Jr, de Oliveira ML: Fumaric aciduria: an overview and the first Brazilian case report. J Inherit Metab Dis. 2010 Aug;33(4):411-9. doi: 10.1007/s10545-010-9134-2. Epub 2010 Jun 15. [PubMed:20549362 ]
| | Obesity |
|---|
- Simone Wahl, Christina Holzapfel, Zhonghao Yu, Michaela Breier, Ivan Kondofersky, Christiane Fuchs, Paula Singmann, Cornelia Prehn, Jerzy Adamski, Harald Grallert, Thomas Illig, Rui Wang-Sattler, Thomas Reinehr (2013). Metabolomics reveals determinants of weight loss during lifestyle intervention in obese children. Metabolomics.
| | Histidinemia |
|---|
- Wevers RA, Engelke U, Wendel U, de Jong JG, Gabreels FJ, Heerschap A: Standardized method for high-resolution 1H-NMR of cerebrospinal fluid. Clin Chem. 1995 May;41(5):744-51. [PubMed:7729054 ]
- Anakura M, Matsuda I, Arashima S, Fukushima N, Oka Y: Histidinemia. Classical and atypical form in siblings. Am J Dis Child. 1975 Jul;129(7):858-61. [PubMed:1146783 ]
- Swarna M, Jyothy A, Usha Rani P, Reddy PP: Amino acid disorders in mental retardation: a two-decade study from Andhra Pradesh. Biochem Genet. 2004 Apr;42(3-4):85-98. [PubMed:15168722 ]
- Nyhan WL, Hilton S: Histidinuria: defective transport of histidine. Am J Med Genet. 1992 Nov 15;44(5):558-61. [PubMed:1481808 ]
- LADU BN, HOWELL RR, JACOBY GA, SEEGMILLER JE, SOBER EK, ZANNONI VG, CANBY JP, ZIEGLER LK: CLINICAL AND BIOCHEMICAL STUDIES ON TWO CASES OF HISTIDINEMIA. Pediatrics. 1963 Aug;32:216-27. [PubMed:14044449 ]
| | Leukemia |
|---|
- Peng CT, Wu KH, Lan SJ, Tsai JJ, Tsai FJ, Tsai CH: Amino acid concentrations in cerebrospinal fluid in children with acute lymphoblastic leukemia undergoing chemotherapy. Eur J Cancer. 2005 May;41(8):1158-63. Epub 2005 Apr 14. [PubMed:15911239 ]
| | Pyridoxamine 5-prime-phosphate oxidase deficiency |
|---|
- Plecko B, Paul K, Paschke E, Stoeckler-Ipsiroglu S, Struys E, Jakobs C, Hartmann H, Luecke T, di Capua M, Korenke C, Hikel C, Reutershahn E, Freilinger M, Baumeister F, Bosch F, Erwa W: Biochemical and molecular characterization of 18 patients with pyridoxine-dependent epilepsy and mutations of the antiquitin (ALDH7A1) gene. Hum Mutat. 2007 Jan;28(1):19-26. [PubMed:17068770 ]
| | Celiac disease |
|---|
- Di Cagno R, De Angelis M, De Pasquale I, Ndagijimana M, Vernocchi P, Ricciuti P, Gagliardi F, Laghi L, Crecchio C, Guerzoni ME, Gobbetti M, Francavilla R: Duodenal and faecal microbiota of celiac children: molecular, phenotype and metabolome characterization. BMC Microbiol. 2011 Oct 4;11:219. doi: 10.1186/1471-2180-11-219. [PubMed:21970810 ]
- De Angelis M, Vannini L, Di Cagno R, Cavallo N, Minervini F, Francavilla R, Ercolini D, Gobbetti M: Salivary and fecal microbiota and metabolome of celiac children under gluten-free diet. Int J Food Microbiol. 2016 Dec 19;239:125-132. doi: 10.1016/j.ijfoodmicro.2016.07.025. Epub 2016 Jul 19. [PubMed:27452636 ]
| | Autism |
|---|
- De Angelis M, Piccolo M, Vannini L, Siragusa S, De Giacomo A, Serrazzanetti DI, Cristofori F, Guerzoni ME, Gobbetti M, Francavilla R: Fecal microbiota and metabolome of children with autism and pervasive developmental disorder not otherwise specified. PLoS One. 2013 Oct 9;8(10):e76993. doi: 10.1371/journal.pone.0076993. eCollection 2013. [PubMed:24130822 ]
| | Crohn's disease |
|---|
- Kolho KL, Pessia A, Jaakkola T, de Vos WM, Velagapudi V: Faecal and Serum Metabolomics in Paediatric Inflammatory Bowel Disease. J Crohns Colitis. 2017 Mar 1;11(3):321-334. doi: 10.1093/ecco-jcc/jjw158. [PubMed:27609529 ]
| | Ulcerative colitis |
|---|
- Kolho KL, Pessia A, Jaakkola T, de Vos WM, Velagapudi V: Faecal and Serum Metabolomics in Paediatric Inflammatory Bowel Disease. J Crohns Colitis. 2017 Mar 1;11(3):321-334. doi: 10.1093/ecco-jcc/jjw158. [PubMed:27609529 ]
| | Perillyl alcohol administration for cancer treatment |
|---|
- Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M: Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics. 2010 Mar;6(1):78-95. Epub 2009 Sep 10. [PubMed:20300169 ]
| | Pancreatic cancer |
|---|
- Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M: Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics. 2010 Mar;6(1):78-95. Epub 2009 Sep 10. [PubMed:20300169 ]
- Zhang L, Jin H, Guo X, Yang Z, Zhao L, Tang S, Mo P, Wu K, Nie Y, Pan Y, Fan D: Distinguishing pancreatic cancer from chronic pancreatitis and healthy individuals by (1)H nuclear magnetic resonance-based metabonomic profiles. Clin Biochem. 2012 Sep;45(13-14):1064-9. doi: 10.1016/j.clinbiochem.2012.05.012. Epub 2012 May 19. [PubMed:22613268 ]
| | Periodontal disease |
|---|
- Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M: Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics. 2010 Mar;6(1):78-95. Epub 2009 Sep 10. [PubMed:20300169 ]
| | Frontotemporal dementia |
|---|
- Tsuruoka M, Hara J, Hirayama A, Sugimoto M, Soga T, Shankle WR, Tomita M: Capillary electrophoresis-mass spectrometry-based metabolome analysis of serum and saliva from neurodegenerative dementia patients. Electrophoresis. 2013 Oct;34(19):2865-72. doi: 10.1002/elps.201300019. Epub 2013 Sep 6. [PubMed:23857558 ]
| | Lewy body disease |
|---|
- Tsuruoka M, Hara J, Hirayama A, Sugimoto M, Soga T, Shankle WR, Tomita M: Capillary electrophoresis-mass spectrometry-based metabolome analysis of serum and saliva from neurodegenerative dementia patients. Electrophoresis. 2013 Oct;34(19):2865-72. doi: 10.1002/elps.201300019. Epub 2013 Sep 6. [PubMed:23857558 ]
| | Autosomal dominant polycystic kidney disease |
|---|
- Gronwald W, Klein MS, Zeltner R, Schulze BD, Reinhold SW, Deutschmann M, Immervoll AK, Boger CA, Banas B, Eckardt KU, Oefner PJ: Detection of autosomal dominant polycystic kidney disease by NMR spectroscopic fingerprinting of urine. Kidney Int. 2011 Jun;79(11):1244-53. doi: 10.1038/ki.2011.30. Epub 2011 Mar 9. [PubMed:21389975 ]
| | Argininosuccinic aciduria |
|---|
- Gronwald W, Klein MS, Kaspar H, Fagerer SR, Nurnberger N, Dettmer K, Bertsch T, Oefner PJ: Urinary metabolite quantification employing 2D NMR spectroscopy. Anal Chem. 2008 Dec 1;80(23):9288-97. doi: 10.1021/ac801627c. [PubMed:19551947 ]
| | Propionic acidemia |
|---|
- Gronwald W, Klein MS, Kaspar H, Fagerer SR, Nurnberger N, Dettmer K, Bertsch T, Oefner PJ: Urinary metabolite quantification employing 2D NMR spectroscopy. Anal Chem. 2008 Dec 1;80(23):9288-97. doi: 10.1021/ac801627c. [PubMed:19551947 ]
| | Tyrosinemia I |
|---|
- Gronwald W, Klein MS, Kaspar H, Fagerer SR, Nurnberger N, Dettmer K, Bertsch T, Oefner PJ: Urinary metabolite quantification employing 2D NMR spectroscopy. Anal Chem. 2008 Dec 1;80(23):9288-97. doi: 10.1021/ac801627c. [PubMed:19551947 ]
| | Phenylketonuria |
|---|
- Gronwald W, Klein MS, Kaspar H, Fagerer SR, Nurnberger N, Dettmer K, Bertsch T, Oefner PJ: Urinary metabolite quantification employing 2D NMR spectroscopy. Anal Chem. 2008 Dec 1;80(23):9288-97. doi: 10.1021/ac801627c. [PubMed:19551947 ]
| | Maple syrup urine disease |
|---|
- Gronwald W, Klein MS, Kaspar H, Fagerer SR, Nurnberger N, Dettmer K, Bertsch T, Oefner PJ: Urinary metabolite quantification employing 2D NMR spectroscopy. Anal Chem. 2008 Dec 1;80(23):9288-97. doi: 10.1021/ac801627c. [PubMed:19551947 ]
| | Eosinophilic esophagitis |
|---|
- Slae, M., Huynh, H., Wishart, D.S. (2014). Analysis of 30 normal pediatric urine samples via NMR spectroscopy (unpublished work). NA.
|
|
|---|
| Associated OMIM IDs | |
|---|
| External Links |
|---|
| DrugBank ID | DB00117 |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FooDB ID | FDB011856 |
|---|
| KNApSAcK ID | C00001363 |
|---|
| Chemspider ID | 6038 |
|---|
| KEGG Compound ID | C00135 |
|---|
| BioCyc ID | HIS |
|---|
| BiGG ID | 33985 |
|---|
| Wikipedia Link | L-histidine |
|---|
| METLIN ID | 21 |
|---|
| PubChem Compound | 6274 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 15971 |
|---|
| Food Biomarker Ontology | Not Available |
|---|
| VMH ID | HIS_L |
|---|
| MarkerDB ID | MDB00000085 |
|---|
| Good Scents ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Aurelio Luigi; Brownlee Robert T C; Hughes Andrew B A novel synthesis of N-methyl asparagine, arginine, histidine, and tryptophan. Organic letters (2002), 4(21), 3767-9. |
|---|
| Material Safety Data Sheet (MSDS) | Not Available |
|---|
| General References | - Xiao B, Jing C, Kelly G, Walker PA, Muskett FW, Frenkiel TA, Martin SR, Sarma K, Reinberg D, Gamblin SJ, Wilson JR: Specificity and mechanism of the histone methyltransferase Pr-Set7. Genes Dev. 2005 Jun 15;19(12):1444-54. Epub 2005 Jun 2. [PubMed:15933069 ]
- Sreekumar A, Poisson LM, Rajendiran TM, Khan AP, Cao Q, Yu J, Laxman B, Mehra R, Lonigro RJ, Li Y, Nyati MK, Ahsan A, Kalyana-Sundaram S, Han B, Cao X, Byun J, Omenn GS, Ghosh D, Pennathur S, Alexander DC, Berger A, Shuster JR, Wei JT, Varambally S, Beecher C, Chinnaiyan AM: Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature. 2009 Feb 12;457(7231):910-4. doi: 10.1038/nature07762. [PubMed:19212411 ]
- Shoemaker JD, Elliott WH: Automated screening of urine samples for carbohydrates, organic and amino acids after treatment with urease. J Chromatogr. 1991 Jan 2;562(1-2):125-38. [PubMed:2026685 ]
- Silwood CJ, Lynch E, Claxson AW, Grootveld MC: 1H and (13)C NMR spectroscopic analysis of human saliva. J Dent Res. 2002 Jun;81(6):422-7. [PubMed:12097436 ]
- Nicholson JK, O'Flynn MP, Sadler PJ, Macleod AF, Juul SM, Sonksen PH: Proton-nuclear-magnetic-resonance studies of serum, plasma and urine from fasting normal and diabetic subjects. Biochem J. 1984 Jan 15;217(2):365-75. [PubMed:6696735 ]
- Wevers RA, Engelke U, Wendel U, de Jong JG, Gabreels FJ, Heerschap A: Standardized method for high-resolution 1H-NMR of cerebrospinal fluid. Clin Chem. 1995 May;41(5):744-51. [PubMed:7729054 ]
- Hagenfeldt L, Bjerkenstedt L, Edman G, Sedvall G, Wiesel FA: Amino acids in plasma and CSF and monoamine metabolites in CSF: interrelationship in healthy subjects. J Neurochem. 1984 Mar;42(3):833-7. [PubMed:6198473 ]
- Peng CT, Wu KH, Lan SJ, Tsai JJ, Tsai FJ, Tsai CH: Amino acid concentrations in cerebrospinal fluid in children with acute lymphoblastic leukemia undergoing chemotherapy. Eur J Cancer. 2005 May;41(8):1158-63. Epub 2005 Apr 14. [PubMed:15911239 ]
- Cynober LA: Plasma amino acid levels with a note on membrane transport: characteristics, regulation, and metabolic significance. Nutrition. 2002 Sep;18(9):761-6. [PubMed:12297216 ]
- Rainesalo S, Keranen T, Palmio J, Peltola J, Oja SS, Saransaari P: Plasma and cerebrospinal fluid amino acids in epileptic patients. Neurochem Res. 2004 Jan;29(1):319-24. [PubMed:14992292 ]
- Klassen P, Furst P, Schulz C, Mazariegos M, Solomons NW: Plasma free amino acid concentrations in healthy Guatemalan adults and in patients with classic dengue. Am J Clin Nutr. 2001 Mar;73(3):647-52. [PubMed:11237944 ]
- Mukerji SK, Pimstone NR, Gandhi SN, Tan KT: Biochemical diagnosis and monitoring therapeutic modulation of disease activity in an unusual case of congenital erythropoietic porphyria. Clin Chem. 1985 Dec;31(12):1946-51. [PubMed:4064282 ]
- Churchill D, Yacoub JM, Siu KP, Symes A, Gault MH: Toxic nephropathy after low-dose methoxyflurane anesthesia: drug interaction with secobarbital? Can Med Assoc J. 1976 Feb 21;114(4):326-8, 333. [PubMed:1253070 ]
- Sieja K, Stanosz S, von Mach-Szczypinski J, Olewniczak S, Stanosz M: Concentration of histamine in serum and tissues of the primary ductal breast cancers in women. Breast. 2005 Jun;14(3):236-41. Epub 2005 Jan 21. [PubMed:15927833 ]
- Masini E, Fabbroni V, Giannini L, Vannacci A, Messerini L, Perna F, Cortesini C, Cianchi F: Histamine and histidine decarboxylase up-regulation in colorectal cancer: correlation with tumor stage. Inflamm Res. 2005 Apr;54 Suppl 1:S80-1. [PubMed:15928846 ]
- Xiao YP, Han CB, Mao XY, Li JY, Xu L, Ren CS, Xin Y: Relationship between abnormality of FHIT gene and EBV infection in gastric cancer. World J Gastroenterol. 2005 Jun 7;11(21):3212-6. [PubMed:15929169 ]
- Mason AB, Halbrooks PJ, James NG, Connolly SA, Larouche JR, Smith VC, MacGillivray RT, Chasteen ND: Mutational analysis of C-lobe ligands of human serum transferrin: insights into the mechanism of iron release. Biochemistry. 2005 Jun 7;44(22):8013-21. [PubMed:15924420 ]
- Janknecht R, Hipskind RA, Houthaeve T, Nordheim A, Stunnenberg HG: Identification of multiple SRF N-terminal phosphorylation sites affecting DNA binding properties. EMBO J. 1992 Mar;11(3):1045-54. [PubMed:1547771 ]
- Peterson JW, Boldogh I, Popov VL, Saini SS, Chopra AK: Anti-inflammatory and antisecretory potential of histidine in Salmonella-challenged mouse small intestine. Lab Invest. 1998 May;78(5):523-34. [PubMed:9605177 ]
- Gerber DA: Low free serum histidine concentration in rheumatoid arthritis. A measure of disease activity. J Clin Invest. 1975 Jun;55(6):1164-73. [PubMed:1079527 ]
- Hemler RJ, Hoogeveen JH, Kraaier V, Van Huffelen AC, Wieneke GH, Hijman R, Glerum JH: A pharmacological model of cerebral ischemia. The effects of indomethacin on cerebral blood flow velocity, quantitative EEG and cognitive functions. Methods Find Exp Clin Pharmacol. 1990 Nov;12(9):641-3. [PubMed:2084459 ]
- Feng RN, Niu YC, Sun XW, Li Q, Zhao C, Wang C, Guo FC, Sun CH, Li Y: Histidine supplementation improves insulin resistance through suppressed inflammation in obese women with the metabolic syndrome: a randomised controlled trial. Diabetologia. 2013 May;56(5):985-94. doi: 10.1007/s00125-013-2839-7. Epub 2013 Jan 30. [PubMed:23361591 ]
- Watanabe M, Suliman ME, Qureshi AR, Garcia-Lopez E, Barany P, Heimburger O, Stenvinkel P, Lindholm B: Consequences of low plasma histidine in chronic kidney disease patients: associations with inflammation, oxidative stress, and mortality. Am J Clin Nutr. 2008 Jun;87(6):1860-6. [PubMed:18541578 ]
- Jung J, Kim SH, Lee HS, Choi GS, Jung YS, Ryu DH, Park HS, Hwang GS: Serum metabolomics reveals pathways and biomarkers associated with asthma pathogenesis. Clin Exp Allergy. 2013 Apr;43(4):425-33. doi: 10.1111/cea.12089. [PubMed:23517038 ]
- Elshenawy S, Pinney SE, Stuart T, Doulias PT, Zura G, Parry S, Elovitz MA, Bennett MJ, Bansal A, Strauss JF 3rd, Ischiropoulos H, Simmons RA: The Metabolomic Signature of the Placenta in Spontaneous Preterm Birth. Int J Mol Sci. 2020 Feb 4;21(3). pii: ijms21031043. doi: 10.3390/ijms21031043. [PubMed:32033212 ]
- Effective method for the amelioration and prevention of tissue and cellular damage [Link]
| Show more...
|---|